The h subunit of eIF3 promotes reinitiation competence during translation of mRNAs harboring upstream open reading frames
- PMID: 20179149
- PMCID: PMC2844622
- DOI: 10.1261/rna.2056010
The h subunit of eIF3 promotes reinitiation competence during translation of mRNAs harboring upstream open reading frames
Abstract
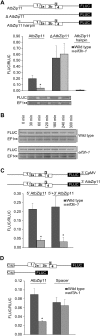
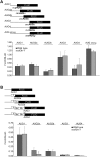
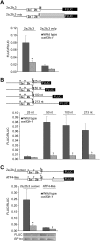
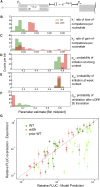
Upstream open reading frames (uORFs) are protein coding elements in the 5' leader of messenger RNAs. uORFs generally inhibit translation of the main ORF because ribosomes that perform translation elongation suffer either permanent or conditional loss of reinitiation competence. After conditional loss, reinitiation competence may be regained by, at the minimum, reacquisition of a fresh methionyl-tRNA. The conserved h subunit of Arabidopsis eukaryotic initiation factor 3 (eIF3) mitigates the inhibitory effects of certain uORFs. Here, we define more precisely how this occurs, by combining gene expression data from mutated 5' leaders of Arabidopsis AtbZip11 (At4g34590) and yeast GCN4 with a computational model of translation initiation in wild-type and eif3h mutant plants. Of the four phylogenetically conserved uORFs in AtbZip11, three are inhibitory to translation, while one is anti-inhibitory. The mutation in eIF3h has no major effect on uORF start codon recognition. Instead, eIF3h supports efficient reinitiation after uORF translation. Modeling suggested that the permanent loss of reinitiation competence during uORF translation occurs at a faster rate in the mutant than in the wild type. Thus, eIF3h ensures that a fraction of uORF-translating ribosomes retain their competence to resume scanning. Experiments using the yeast GCN4 leader provided no evidence that eIF3h fosters tRNA reaquisition. Together, these results attribute a specific molecular function in translation initiation to an individual eIF3 subunit in a multicellular eukaryote.
Figures



Similar articles
-
Translation reinitiation and development are compromised in similar ways by mutations in translation initiation factor eIF3h and the ribosomal protein RPL24.BMC Plant Biol. 2010 Aug 27;10:193. doi: 10.1186/1471-2229-10-193. BMC Plant Biol. 2010. PMID: 20799971 Free PMC article.
-
Fail-safe mechanism of GCN4 translational control--uORF2 promotes reinitiation by analogous mechanism to uORF1 and thus secures its key role in GCN4 expression.Nucleic Acids Res. 2014 May;42(9):5880-93. doi: 10.1093/nar/gku204. Epub 2014 Mar 12. Nucleic Acids Res. 2014. PMID: 24623812 Free PMC article.
-
Translational regulation via 5' mRNA leader sequences revealed by mutational analysis of the Arabidopsis translation initiation factor subunit eIF3h.Plant Cell. 2004 Dec;16(12):3341-56. doi: 10.1105/tpc.104.026880. Epub 2004 Nov 17. Plant Cell. 2004. PMID: 15548739 Free PMC article.
-
Does eIF3 promote reinitiation after translation of short upstream ORFs also in mammalian cells?RNA Biol. 2017 Dec 2;14(12):1660-1667. doi: 10.1080/15476286.2017.1353863. Epub 2017 Sep 15. RNA Biol. 2017. PMID: 28745933 Free PMC article. Review.
-
Translational regulation by uORFs and start codon selection stringency.Genes Dev. 2023 Jun 1;37(11-12):474-489. doi: 10.1101/gad.350752.123. Epub 2023 Jul 11. Genes Dev. 2023. PMID: 37433636 Free PMC article. Review.
Cited by
-
Arabidopsis ribosomal proteins control developmental programs through translational regulation of auxin response factors.Proc Natl Acad Sci U S A. 2012 Nov 27;109(48):19537-44. doi: 10.1073/pnas.1214774109. Epub 2012 Nov 9. Proc Natl Acad Sci U S A. 2012. PMID: 23144218 Free PMC article.
-
Interaction of PABPC1 with the translation initiation complex is critical to the NMD resistance of AUG-proximal nonsense mutations.Nucleic Acids Res. 2012 Feb;40(3):1160-73. doi: 10.1093/nar/gkr820. Epub 2011 Oct 11. Nucleic Acids Res. 2012. PMID: 21989405 Free PMC article.
-
RNA granules: the good, the bad and the ugly.Cell Signal. 2011 Feb;23(2):324-34. doi: 10.1016/j.cellsig.2010.08.011. Epub 2010 Sep 8. Cell Signal. 2011. PMID: 20813183 Free PMC article. Review.
-
Pharmacological dimerization and activation of the exchange factor eIF2B antagonizes the integrated stress response.Elife. 2015 Apr 15;4:e07314. doi: 10.7554/eLife.07314. Elife. 2015. PMID: 25875391 Free PMC article.
-
The impact of the phosphomimetic eIF2αS/D on global translation, reinitiation and the integrated stress response is attenuated in N2a cells.Nucleic Acids Res. 2015 Sep 30;43(17):8392-404. doi: 10.1093/nar/gkv827. Epub 2015 Aug 11. Nucleic Acids Res. 2015. PMID: 26264663 Free PMC article.
References
-
- Abastado JP, Miller PF, Hinnebusch AG. A quantitative model for translational control of the GCN4 gene of Saccharomyces cerevisiae. New Biol. 1991a;3:511–524. - PubMed
-
- Angenon G, Van Montagu M, Depicker A. Analysis of the stop codon context in plant nuclear genes. FEBS Lett. 1990;271:144–146. - PubMed
-
- Ashlock D. Evolutionary computation for modeling and optimization. Springer; New York: 2006.
-
- Benne R, Hershey JW. The mechanism of action of protein synthesis initiation factors from rabbit reticulocytes. J Biol Chem. 1978;253:3078–3087. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
