Arginylation regulates intracellular actin polymer level by modulating actin properties and binding of capping and severing proteins
- PMID: 20181827
- PMCID: PMC2854093
- DOI: 10.1091/mbc.e09-09-0829
Arginylation regulates intracellular actin polymer level by modulating actin properties and binding of capping and severing proteins
Abstract
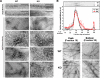
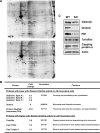
Actin arginylation regulates lamella formation in motile fibroblasts, but the underlying molecular mechanisms are unknown. To understand how arginylation affects the actin cytoskeleton, we investigated the biochemical properties and the structural organization of actin filaments in wild-type and arginyltransferase (Ate1) knockout cells. We found that Ate1 knockout results in a dramatic reduction of the actin polymer levels in vivo accompanied by a corresponding increase in the monomer level. Purified nonarginylated actin has altered polymerization properties, and actin filaments from Ate1 knockout cells show altered interactions with several associated proteins. Ate1 knockout cells have severe impairment of cytoskeletal organization throughout the cell. Thus, arginylation regulates the ability of actin to form filaments in the whole cell rather than preventing the collapse of preformed actin networks at the cell leading edge as proposed in our previous model. This regulation is achieved through interconnected mechanisms that involve actin polymerization per se and through binding of actin-associated proteins.
Figures






Similar articles
-
Arginylation of beta-actin regulates actin cytoskeleton and cell motility.Science. 2006 Jul 14;313(5784):192-6. doi: 10.1126/science.1129344. Epub 2006 Jun 22. Science. 2006. PMID: 16794040
-
Arginyltransferase regulates alpha cardiac actin function, myofibril formation and contractility during heart development.Development. 2008 Dec;135(23):3881-9. doi: 10.1242/dev.022723. Epub 2008 Oct 23. Development. 2008. PMID: 18948421 Free PMC article.
-
Distinct roles of four gelsolin-like domains of Caenorhabditis elegans gelsolin-like protein-1 in actin filament severing, barbed end capping, and phosphoinositide binding.Biochemistry. 2010 May 25;49(20):4349-60. doi: 10.1021/bi100215b. Biochemistry. 2010. PMID: 20392036 Free PMC article.
-
Protein arginylation, a global biological regulator that targets actin cytoskeleton and the muscle.Anat Rec (Hoboken). 2014 Sep;297(9):1630-6. doi: 10.1002/ar.22969. Anat Rec (Hoboken). 2014. PMID: 25125176 Free PMC article. Review.
-
From filopodia to synapses: the role of actin-capping and anti-capping proteins.Eur J Neurosci. 2011 Nov;34(10):1655-62. doi: 10.1111/j.1460-9568.2011.07897.x. Eur J Neurosci. 2011. PMID: 22103422 Review.
Cited by
-
Protein Arginylation: Milestones of Discovery.Methods Mol Biol. 2023;2620:1-13. doi: 10.1007/978-1-0716-2942-0_1. Methods Mol Biol. 2023. PMID: 37010742
-
Correlated Measurement of Endogenous ATE1 Activity on Native Acceptor Proteins in Tissues and Cultured Cells to Detect Cellular Aging.Methods Mol Biol. 2023;2620:41-50. doi: 10.1007/978-1-0716-2942-0_6. Methods Mol Biol. 2023. PMID: 37010747
-
PHD3-mediated prolyl hydroxylation of nonmuscle actin impairs polymerization and cell motility.Mol Biol Cell. 2014 Sep 15;25(18):2788-96. doi: 10.1091/mbc.E14-02-0775. Epub 2014 Jul 30. Mol Biol Cell. 2014. PMID: 25079693 Free PMC article.
-
Modular actin nano-architecture enables podosome protrusion and mechanosensing.Nat Commun. 2019 Nov 15;10(1):5171. doi: 10.1038/s41467-019-13123-3. Nat Commun. 2019. PMID: 31729386 Free PMC article.
-
Posttranslational arginylation as a global biological regulator.Dev Biol. 2011 Oct 1;358(1):1-8. doi: 10.1016/j.ydbio.2011.06.043. Epub 2011 Jul 18. Dev Biol. 2011. PMID: 21784066 Free PMC article. Review.
References
-
- Balzi E., Choder M., Chen W. N., Varshavsky A., Goffeau A. Cloning and functional analysis of the arginyl-tRNA-protein transferase gene ATE1 of Saccharomyces cerevisiae. J. Biol. Chem. 1990;265:7464–7471. - PubMed
-
- Burtnick L. D., Koepf E. K., Grimes J., Jones E. Y., Stuart D. I., McLaughlin P. J., Robinson R. C. The crystal structure of plasma gelsolin: implications for actin severing, capping, and nucleation. Cell. 1997;90:661–670. - PubMed
-
- Carlier M. F., Pantaloni D. Control of actin dynamics in cell motility. J. Mol. Biol. 1997;269:459–467. - PubMed
-
- Cooper J. A., Pollard T. D. Methods to measure actin polymerization. Methods Enzymol. 1982;85(Pt B):182–210. - PubMed
-
- Cooper J. A., Schafer D. A. Control of actin assembly and disassembly at filament ends. Curr. Opin. Cell Biol. 2000;12:97–103. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

