Hearts of hypoxia-inducible factor prolyl 4-hydroxylase-2 hypomorphic mice show protection against acute ischemia-reperfusion injury
- PMID: 20185832
- PMCID: PMC2859527
- DOI: 10.1074/jbc.M109.084855
Hearts of hypoxia-inducible factor prolyl 4-hydroxylase-2 hypomorphic mice show protection against acute ischemia-reperfusion injury
Abstract
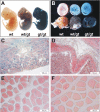
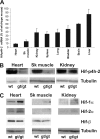
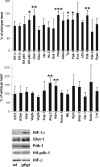
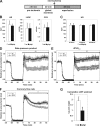
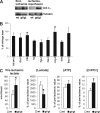
Hypoxia-inducible factor (HIF) has a pivotal role in oxygen homeostasis and cardioprotection mediated by ischemic preconditioning. Its stability is regulated by HIF prolyl 4-hydroxylases (HIF-P4Hs), the inhibition of which is regarded as a promising strategy for treating diseases such as anemia and ischemia. We generated a viable Hif-p4h-2 hypomorph mouse line (Hif-p4h-2(gt/gt)) that expresses decreased amounts of wild-type Hif-p4h-2 mRNA: 8% in the heart; 15% in the skeletal muscle; 34-47% in the kidney, spleen, lung, and bladder; 60% in the brain; and 85% in the liver. These mice have no polycythemia and show no signs of the dilated cardiomyopathy or hyperactive angiogenesis observed in mice with broad spectrum conditional Hif-p4h-2 inactivation. We focused here on the effects of chronic Hif-p4h-2 deficiency in the heart. Hif-1 and Hif-2 were stabilized, and the mRNA levels of glucose transporter-1, several enzymes of glycolysis, pyruvate dehydrogenase kinase 1, angiopoietin-2, and adrenomedullin were increased in the Hif-p4h-2(gt/gt) hearts. When isolated Hif-p4h-2(gt/gt) hearts were subjected to ischemia-reperfusion, the recovery of mechanical function and coronary flow rate was significantly better than in wild type, while cumulative release of lactate dehydrogenase reflecting the infarct size was reduced. The preischemic amount of lactate was increased, and the ischemic versus preischemic [CrP]/[Cr] and [ATP] remained at higher levels in Hif-p4h-2(gt/gt) hearts, indicating enhanced glycolysis and an improved cellular energy state. Our data suggest that chronic stabilization of Hif-1alpha and Hif-2alpha by genetic knockdown of Hif-p4h-2 promotes cardioprotection by induction of many genes involved in glucose metabolism, cardiac function, and blood pressure.
Figures
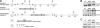
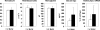

References
-
- Kaelin W. G., Jr., Ratcliffe P. J. (2008) Mol. Cell 30, 393–402 - PubMed
-
- Fraisl P., Aragonés J., Carmeliet P. (2009) Nat. Rev. Drug Discov. 8, 139–152 - PubMed
-
- Semenza G. L. (2009) Physiology 24, 97–106 - PubMed
-
- Ivan M., Kondo K., Yang H., Kim W., Valiando J., Ohh M., Salic A., Asara J. M., Lane W. S., Kaelin W. G., Jr. (2001) Science 292, 464–468 - PubMed
-
- Jaakkola P., Mole D. R., Tian Y. M., Wilson M. I., Gielbert J., Gaskell S. J., Kriegsheim Av, Hebestreit H. F., Mukherji M., Schofield C. J., Maxwell P. H., Pugh C. W., Ratcliffe P. J. (2001) Science 292, 468–472 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

