A computational approach for deciphering the organization of glycosaminoglycans
- PMID: 20186334
- PMCID: PMC2826411
- DOI: 10.1371/journal.pone.0009389
A computational approach for deciphering the organization of glycosaminoglycans
Abstract
Background: Increasing evidence has revealed important roles for complex glycans as mediators of normal and pathological processes. Glycosaminoglycans are a class of glycans that bind and regulate the function of a wide array of proteins at the cell-extracellular matrix interface. The specific sequence and chemical organization of these polymers likely define function; however, identification of the structure-function relationships of glycosaminoglycans has been met with challenges associated with the unique level of complexity and the nontemplate-driven biosynthesis of these biopolymers.
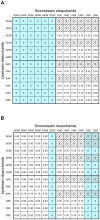
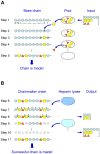
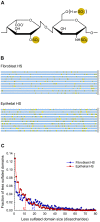
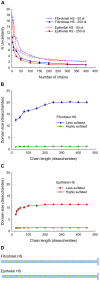
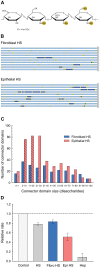
Methodology/principal findings: To address these challenges, we have devised a computational approach to predict fine structure and patterns of domain organization of the specific glycosaminoglycan, heparan sulfate (HS). Using chemical composition data obtained after complete and partial digestion of mixtures of HS chains with specific degradative enzymes, the computational analysis produces populations of theoretical HS chains with structures that meet both biosynthesis and enzyme degradation rules. The model performs these operations through a modular format consisting of input/output sections and three routines called chainmaker, chainbreaker, and chainsorter. We applied this methodology to analyze HS preparations isolated from pulmonary fibroblasts and epithelial cells. Significant differences in the general organization of these two HS preparations were observed, with HS from epithelial cells having a greater frequency of highly sulfated domains. Epithelial HS also showed a higher density of specific HS domains that have been associated with inhibition of neutrophil elastase. Experimental analysis of elastase inhibition was consistent with the model predictions and demonstrated that HS from epithelial cells had greater inhibitory activity than HS from fibroblasts.
Conclusions/significance: This model establishes the conceptual framework for a new class of computational tools to use to assess patterns of domain organization within glycosaminoglycans. These tools will provide a means to consider high-level chain organization in deciphering the structure-function relationships of polysaccharides in biology.
Conflict of interest statement
Figures
References
-
- Bernfield M, Gotte M, Park PW, Reizes O, Fitzgerald ML, et al. Functions of cell surface heparan sulfate proteoglycans. Annu Rev Biochem. 1999;68:729–777. - PubMed
-
- Gandhi NS, Mancera RL. The structure of glycosaminoglycans and their interactions with proteins. Chem Biol Drug Des. 2008;72:455–482. - PubMed
-
- Lindahl U. Heparan sulfate-protein interactions–a concept for drug design? Thromb Haemost. 2007;98:109–115. - PubMed
-
- Raman R, Raguram S, Venkataraman G, Paulson JC, Sasisekharan R. Glycomics: an integrated systems approach to structure-function relationships of glycans. Nat Methods. 2005;2:817–824. - PubMed
-
- Sasisekharan R, Raman R, Prabhakar V. Glycomics approach to structure-function relationships of glycosaminoglycans. Annu Rev Biomed Eng. 2006;8:181–231. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

