Helix-sheet packing in proteins
- PMID: 20186972
- PMCID: PMC2854864
- DOI: 10.1002/prot.22688
Helix-sheet packing in proteins
Abstract
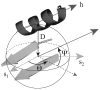
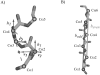
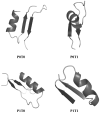
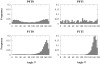
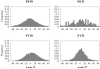
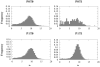
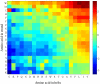
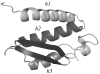
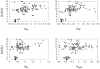
The three-dimensional structure of a protein is organized around the packing of its secondary structure elements. Although much is known about the packing geometry observed between alpha-helices and between beta-sheets, there has been little progress on characterizing helix-sheet interactions. We present an analysis of the conformation of alphabeta(2) motifs in proteins, corresponding to all occurrences of helices in contact with two strands that are hydrogen bonded. The geometry of the alphabeta(2) motif is characterized by the azimuthal angle theta between the helix axis and an average vector representing the two strands, the elevation angle psi between the helix axis and the plane containing the two strands, and the distance D between the helix and the strands. We observe that the helix tends to align to the two strands, with a preference for an antiparallel orientation if the two strands are parallel; this preference is diminished for other topologies of the beta-sheet. Side-chain packing at the interface between the helix and the strands is mostly hydrophobic, with a preference for aliphatic amino acids in the strand and aromatic amino acids in the helix. From the knowledge of the geometry and amino acid propensities of alphabeta(2) motifs in proteins, we have derived different statistical potentials that are shown to be efficient in picking native-like conformations among a set of non-native conformations in well-known decoy datasets. The information on the geometry of alphabeta(2) motifs as well as the related statistical potentials have applications in the field of protein structure prediction.
Proteins 2010. (c) 2010 Wiley-Liss, Inc.
Figures
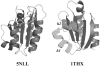
Similar articles
-
PackHelix: a tool for helix-sheet packing during protein structure prediction.Proteins. 2011 Oct;79(10):2828-43. doi: 10.1002/prot.23108. Epub 2011 Aug 23. Proteins. 2011. PMID: 21905109 Free PMC article.
-
Discovery of a significant, nontopological preference for antiparallel alignment of helices with parallel regions in sheets.Protein Sci. 2003 May;12(5):1119-25. doi: 10.1110/ps.0238803. Protein Sci. 2003. PMID: 12717033 Free PMC article.
-
Interactions between an alpha-helix and a beta-sheet. Energetics of alpha/beta packing in proteins.J Mol Biol. 1985 Dec 5;186(3):591-609. doi: 10.1016/0022-2836(85)90133-0. J Mol Biol. 1985. PMID: 4093981
-
Parallel and antiparallel β-strands differ in amino acid composition and availability of short constituent sequences.J Chem Inf Model. 2011 Jun 27;51(6):1457-64. doi: 10.1021/ci200027d. Epub 2011 May 11. J Chem Inf Model. 2011. PMID: 21520893
-
[A turning point in the knowledge of the structure-function-activity relations of elastin].J Soc Biol. 2001;195(2):181-93. J Soc Biol. 2001. PMID: 11727705 Review. French.
Cited by
-
An amino acid code to define a protein's tertiary packing surface.Proteins. 2016 Feb;84(2):201-16. doi: 10.1002/prot.24966. Epub 2015 Dec 22. Proteins. 2016. PMID: 26575337 Free PMC article.
-
PackHelix: a tool for helix-sheet packing during protein structure prediction.Proteins. 2011 Oct;79(10):2828-43. doi: 10.1002/prot.23108. Epub 2011 Aug 23. Proteins. 2011. PMID: 21905109 Free PMC article.
-
Description of local and global shape properties of protein helices.J Mol Model. 2013 Jul;19(7):2901-11. doi: 10.1007/s00894-013-1819-7. Epub 2013 Mar 27. J Mol Model. 2013. PMID: 23529181
-
An amino acid packing code for α-helical structure and protein design.J Mol Biol. 2012 Jun 8;419(3-4):234-54. doi: 10.1016/j.jmb.2012.03.004. Epub 2012 Mar 15. J Mol Biol. 2012. PMID: 22426125 Free PMC article.
-
Switching an active site helix in dihydrofolate reductase reveals limits to subdomain modularity.Biophys J. 2021 Nov 2;120(21):4738-4750. doi: 10.1016/j.bpj.2021.09.032. Epub 2021 Sep 25. Biophys J. 2021. PMID: 34571014 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

