Identification of functional modules that correlate with phenotypic difference: the influence of network topology
- PMID: 20187943
- PMCID: PMC2872883
- DOI: 10.1186/gb-2010-11-2-r23
Identification of functional modules that correlate with phenotypic difference: the influence of network topology
Abstract
One of the important challenges to post-genomic biology is relating observed phenotypic alterations to the underlying collective alterations in genes. Current inferential methods, however, invariably omit large bodies of information on the relationships between genes. We present a method that takes account of such information - expressed in terms of the topology of a correlation network - and we apply the method in the context of current procedures for gene set enrichment analysis.
Figures
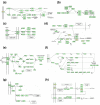



References
-
- Goeman JJ, Buhlmann P. Analyzing gene expression data in terms of gene sets: methodological issues. Bioinformatics. 2007;23:980–987. - PubMed
-
- Pavlidis P, Lewis DP, Noble WS. Exploring gene expression data with class scores. Pac Symp Biocomput. 2002. pp. 474–485. - PubMed
-
- Mootha VK, Lindgren CM, Eriksson KF, Subramanian A, Sihag S, Lehar J, Puigserver P, Carlsson E, Ridderstrale M, Laurila E, Houstis N, Daly MJ, Patterson N, Mesirov JP, Golub TR, Tamayo P, Spiegelman B, Lander ES, Hirschhorn JN, Altshuler D, Groop LC. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet. 2003;34:267–273. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

