Neuronal MeCP2 is expressed at near histone-octamer levels and globally alters the chromatin state
- PMID: 20188665
- PMCID: PMC4338610
- DOI: 10.1016/j.molcel.2010.01.030
Neuronal MeCP2 is expressed at near histone-octamer levels and globally alters the chromatin state
Abstract
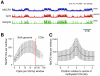
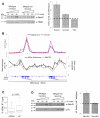
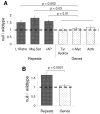
MeCP2 is a nuclear protein with an affinity for methylated DNA that can recruit histone deacetylases. Deficiency or excess of MeCP2 causes severe neurological problems, suggesting that the number of molecules per cell must be precisely regulated. We quantified MeCP2 in neuronal nuclei and found that it is nearly as abundant as the histone octamer. Despite this high abundance, MeCP2 associates preferentially with methylated regions, and high-throughput sequencing showed that its genome-wide binding tracks methyl-CpG density. MeCP2 deficiency results in global changes in neuronal chromatin structure, including elevated histone acetylation and a doubling of histone H1. Neither change is detectable in glia, where MeCP2 occurs at lower levels. The mutant brain also shows elevated transcription of repetitive elements. Our data argue that MeCP2 may not act as a gene-specific transcriptional repressor in neurons, but might instead dampen transcriptional noise genome-wide in a DNA methylation-dependent manner.
Copyright 2010 Elsevier Inc. All rights reserved.
Figures



Comment in
-
A bird's-eye view of MeCP2 binding.Mol Cell. 2010 Feb 26;37(4):451-2. doi: 10.1016/j.molcel.2010.02.006. Mol Cell. 2010. PMID: 20188662
References
-
- Amir RE, Van den Veyver IB, Wan M, Tran CQ, Francke U, Zoghbi HY. Rett syndrome is caused by mutations in X-linked MECP2, encoding methyl-CpG-binding protein 2. Nat Genet. 1999;23:185–188. - PubMed
-
- Armstrong DD. Neuropathology of Rett syndrome. Ment Retard Dev Disabil Res Rev. 2002;8:72–76. - PubMed
-
- Bading H. Transcription-dependent neuronal plasticity the nuclear calcium hypothesis. Eur J Biochem. 2000;267:5280–5283. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

