Tailoring the switch from IRES-dependent to 5'-end-dependent translation with the RNase P ribozyme
- PMID: 20194518
- PMCID: PMC2844631
- DOI: 10.1261/rna.1973710
Tailoring the switch from IRES-dependent to 5'-end-dependent translation with the RNase P ribozyme
Abstract
Translation initiation driven by internal ribosome entry site (IRES) elements is dependent on the structural organization of the IRES region. We have previously shown that a structural motif within the foot-and-mouth-disease virus IRES is recognized in vitro as substrate for the Synechocystis sp. RNase P ribozyme. Here we show that this structure-dependent endonuclease recognizes the IRES element in cultured cells, leading to inhibition of translation. Inhibition of IRES activity was dependent on the expression of the active ribozyme RNA subunit. Moreover, expression of the antisense sequence of the ribozyme did not inhibit IRES activity, demonstrating that stable RNA structures located upstream of the IRES element do not interfere with internal initiation. RNAs carrying defective IRES mutants that were substrates of the ribozyme in vivo revealed an increased translation of the reporter in response to the expression of the active ribozyme. In support of RNA cleavage, subsequent analysis of the translation initiation manner indicated a switch from IRES-dependent to 5'-end-dependent translation of RNase P target RNAs. We conclude that the IRES element is inactivated by expression in cis of RNase P in the cytoplasm of cultured cells, providing a promising antiviral tool to combat picornavirus infections. Furthermore, our results reinforce the essential role of the structural motif that serves as RNase P recognition motif for IRES activity.
Figures






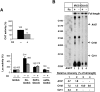
Similar articles
-
Characterization of a cyanobacterial RNase P ribozyme recognition motif in the IRES of foot-and-mouth disease virus reveals a unique structural element.RNA. 2007 Jun;13(6):849-59. doi: 10.1261/rna.506607. Epub 2007 Apr 20. RNA. 2007. PMID: 17449727 Free PMC article.
-
Involvement of proteasome alpha-subunit PSMA7 in hepatitis C virus internal ribosome entry site-mediated translation.Mol Cell Biol. 2001 Dec;21(24):8357-64. doi: 10.1128/MCB.21.24.8357-8364.2001. Mol Cell Biol. 2001. PMID: 11713272 Free PMC article.
-
[Determination of the in vitro antiviral activity of an engineered M1GS ribozyme that targets to the core gene of hepatitis C virus].Wei Sheng Wu Xue Bao. 2013 Aug 4;53(8):875-81. Wei Sheng Wu Xue Bao. 2013. PMID: 24341280 Chinese.
-
Developing RNase P ribozymes for gene-targeting and antiviral therapy.Cell Microbiol. 2004 Jun;6(6):499-508. doi: 10.1111/j.1462-5822.2004.00398.x. Cell Microbiol. 2004. PMID: 15104592 Review.
-
Engineering of RNase P Ribozymes for Therapy against Human Cytomegalovirus Infection.Viruses. 2024 Jul 25;16(8):1196. doi: 10.3390/v16081196. Viruses. 2024. PMID: 39205170 Free PMC article. Review.
Cited by
-
Role of RNA structure motifs in IRES-dependent translation initiation of the coxsackievirus B3: new insights for developing live-attenuated strains for vaccines and gene therapy.Mol Biotechnol. 2013 Oct;55(2):179-202. doi: 10.1007/s12033-013-9674-4. Mol Biotechnol. 2013. PMID: 23881360 Review.
-
RNA structural elements of hepatitis C virus controlling viral RNA translation and the implications for viral pathogenesis.Viruses. 2012 Oct 19;4(10):2233-50. doi: 10.3390/v4102233. Viruses. 2012. PMID: 23202462 Free PMC article. Review.
-
Alternative Mechanisms to Initiate Translation in Eukaryotic mRNAs.Comp Funct Genomics. 2012;2012:391546. doi: 10.1155/2012/391546. Epub 2012 Feb 16. Comp Funct Genomics. 2012. PMID: 22536116 Free PMC article.
References
-
- Alifano P, Rivellini F, Piscitelli C, Arraiano CM, Bruni CB, Carlomagno MS. Ribonuclease E provides substrates for ribonuclease P-dependent processing of a polycistronic mRNA. Genes & Dev. 1994;8:3021–3031. - PubMed
-
- Belsham GJ. Divergent picornavirus IRES elements. Virus Res. 2009;139:183–192. - PubMed
-
- Brannvall M, Kikovska E, Wu S, Kirsebom LA. Evidence for induced fit in bacterial RNase P RNA-mediated cleavage. J Mol Biol. 2007;372:1149–1164. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
