Strong intranucleoid interactions organize the Escherichia coli chromosome into a nucleoid filament
- PMID: 20194778
- PMCID: PMC2841910
- DOI: 10.1073/pnas.0912062107
Strong intranucleoid interactions organize the Escherichia coli chromosome into a nucleoid filament
Abstract
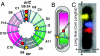
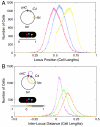
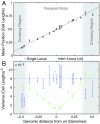
The stochasticity of chromosome organization was investigated by fluorescently labeling genetic loci in live Escherichia coli cells. In spite of the common assumption that the chromosome is well modeled by an unstructured polymer, measurements of the locus distributions reveal that the E. coli chromosome is precisely organized into a nucleoid filament with a linear order. Loci in the body of the nucleoid show a precision of positioning within the cell of better than 10% of the cell length. The precision of interlocus distance of genomically-proximate loci was better than 4% of the cell length. The measured dependence of the precision of interlocus distance on genomic distance singles out intranucleoid interactions as the mechanism responsible for chromosome organization. From the magnitude of the variance, we infer the existence of an as-yet uncharacterized higher-order DNA organization in bacteria. We demonstrate that both the stochastic and average structure of the nucleoid is captured by a fluctuating elastic filament model.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

