Roles of arrest-defective protein 1(225) and hypoxia-inducible factor 1alpha in tumor growth and metastasis
- PMID: 20194889
- PMCID: PMC2841038
- DOI: 10.1093/jnci/djq026
Roles of arrest-defective protein 1(225) and hypoxia-inducible factor 1alpha in tumor growth and metastasis
Abstract
Background: Vascular endothelial growth factor A (VEGFA), a critical mediator of tumor angiogenesis, is a well-characterized target of hypoxia-inducible factor 1 (HIF-1). Murine arrest-defective protein 1A (mARD1A(225)) acetylates HIF-1alpha, triggering its degradation, and thus may play a role in decreased expression of VEGFA.
Methods: We generated Apc(Min/+)/mARD1A(225) transgenic mice and quantified growth of intestinal polyps. Human gastric MKN74 and murine melanoma B16F10 cells overexpressing mARD1A(225) were injected into mice, and tumor growth and metastasis were measured. VEGFA expression and microvessel density in tumors were assessed using immunohistochemistry. To evaluate the role of mARD1A(225) acetylation of Lys532 in HIF-1alpha, we injected B16F10-mARD1A(225) cell lines stably expressing mutant HIF-1alpha/K532R into mice and measured metastasis. All statistical tests were two-sided, and P values less than .05 were considered statistically significant.
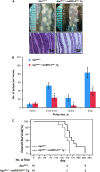
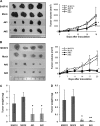
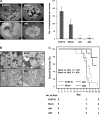
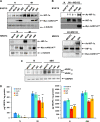
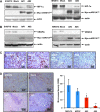
Results: Apc(Min/+)/mARD1A(225) transgenic mice (n = 25) had statistically significantly fewer intestinal polyps than Apc(Min/+) mice (n = 21) (number of intestinal polyps per mouse: Apc(Min/+) mice vs Apc(Min/+)/mARD1A(225) transgenic mice, mean = 83.4 vs 38.0 polyps, difference = 45.4 polyps, 95% confidence interval [CI] = 41.8 to 48.6; P < .001). The growth and metastases of transplanted tumors were also statistically significantly reduced in mice injected with mARD1A(225)-overexpressing cells than in mice injected with control cells (P < .01). Moreover, overexpression of mARD1A(225) decreased VEGFA expression and microvessel density in tumor xenografts (P < .04) and Apc(Min/+) intestinal polyps (P = .001). Mutation of lysine 532 of HIF-1alpha in B16F10-mARD1A(225) cells prevented HIF-1alpha degradation and inhibited the antimetastatic effect of mARD1A(225) (P < .001).
Conclusion: mARD1A(225) may be a novel upstream target that blocks VEGFA expression and tumor-related angiogenesis.
Figures





Similar articles
-
Effect of connective tissue growth factor on hypoxia-inducible factor 1alpha degradation and tumor angiogenesis.J Natl Cancer Inst. 2006 Jul 19;98(14):984-95. doi: 10.1093/jnci/djj242. J Natl Cancer Inst. 2006. PMID: 16849681
-
Role of hypoxia-inducible factor 1alpha in gastric cancer cell growth, angiogenesis, and vessel maturation.J Natl Cancer Inst. 2004 Jun 16;96(12):946-56. doi: 10.1093/jnci/djh168. J Natl Cancer Inst. 2004. PMID: 15199114
-
A hypoxia-dependent upregulation of hypoxia-inducible factor-1 by nuclear factor-κB promotes gastric tumour growth and angiogenesis.Br J Cancer. 2011 Jan 4;104(1):166-74. doi: 10.1038/sj.bjc.6606020. Epub 2010 Nov 30. Br J Cancer. 2011. PMID: 21119667 Free PMC article.
-
BH4 domain of bcl-2 protein is required for its proangiogenic function under hypoxic condition.Carcinogenesis. 2013 Nov;34(11):2558-67. doi: 10.1093/carcin/bgt242. Epub 2013 Jul 8. Carcinogenesis. 2013. PMID: 23836782
-
Progress and perspectives on BMP9-ID1 activation of HIF-1α and VEGFA to promote angiogenesis in hepatic alveolar echinococcosis.Front Oncol. 2024 Nov 15;14:1480683. doi: 10.3389/fonc.2024.1480683. eCollection 2024. Front Oncol. 2024. PMID: 39619441 Free PMC article. Review.
Cited by
-
Nuclear translocation of hARD1 contributes to proper cell cycle progression.PLoS One. 2014 Aug 18;9(8):e105185. doi: 10.1371/journal.pone.0105185. eCollection 2014. PLoS One. 2014. PMID: 25133627 Free PMC article.
-
Energy metabolism in H460 lung cancer cells: effects of histone deacetylase inhibitors.PLoS One. 2011;6(7):e22264. doi: 10.1371/journal.pone.0022264. Epub 2011 Jul 18. PLoS One. 2011. PMID: 21789245 Free PMC article.
-
ARD1/NAA10 in hepatocellular carcinoma: pathways and clinical implications.Exp Mol Med. 2018 Jul 27;50(7):1-12. doi: 10.1038/s12276-018-0106-1. Exp Mol Med. 2018. PMID: 30054466 Free PMC article. Review.
-
High expression of Naa10p associates with lymph node metastasis and predicts favorable prognosis of oral squamous cell carcinoma.Tumour Biol. 2016 May;37(5):6719-28. doi: 10.1007/s13277-015-4563-z. Epub 2015 Dec 9. Tumour Biol. 2016. PMID: 26662107
-
N-α-acetyltransferase 10 (NAA10) in development: the role of NAA10.Exp Mol Med. 2018 Jul 27;50(7):1-11. doi: 10.1038/s12276-018-0105-2. Exp Mol Med. 2018. PMID: 30054454 Free PMC article. Review.
References
-
- Whiteway M, Szostak JW. The ARD1 gene of yeast functions in the switch between the mitotic cell cycle and alternative developmental pathways. Cell. 1985;43(2, pt 1):483–492. - PubMed
-
- Jeong JW, Bae MK, Ahn MY, et al. Regulation and destabilization of HIF-1alpha by ARD1-mediated acetylation. Cell. 2002;111(5):709–720. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

