Visualization of macromolecular structures
- PMID: 20195256
- PMCID: PMC7097155
- DOI: 10.1038/nmeth.1427
Visualization of macromolecular structures
Abstract
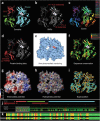
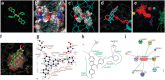
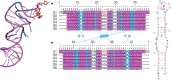
Structural biology is rapidly accumulating a wealth of detailed information about protein function, binding sites, RNA, large assemblies and molecular motions. These data are increasingly of interest to a broader community of life scientists, not just structural experts. Visualization is a primary means for accessing and using these data, yet visualization is also a stumbling block that prevents many life scientists from benefiting from three-dimensional structural data. In this review, we focus on key biological questions where visualizing three-dimensional structures can provide insight and describe available methods and tools.
Conflict of interest statement
The authors declare no competing financial interests.
Figures

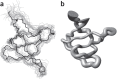
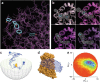
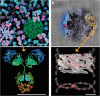

References
-
- Berman H, Henrick K, Nakamura H. Announcing the worldwide Protein Data Bank. Nat. Struct. Biol. 2003;10:980. - PubMed
-
- Tate J. Molecular visualization. Methods Biochem. Anal. 2003;44:135–158. - PubMed
-
- Olson AJ, Pique ME. Visualizing the future of molecular graphics. SAR QSAR Environ. Res. 1998;8:233–247. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

