Modified intracellular-associated phenotypes in a recombinant Salmonella Typhi expressing S. Typhimurium SPI-3 sequences
- PMID: 20195364
- PMCID: PMC2827545
- DOI: 10.1371/journal.pone.0009394
Modified intracellular-associated phenotypes in a recombinant Salmonella Typhi expressing S. Typhimurium SPI-3 sequences
Abstract
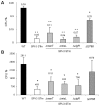
A bioinformatics comparison of Salmonella Pathogenicity Island 3 sequences from S. Typhi and S. Typhimurium serovars showed that ten genes are highly conserved. However three of them are pseudogenes in S. Typhi. Our aim was to understand what functions are lost in S. Typhi due to pseudogenes by constructing a S. Typhi genetic hybrid carrying the SPI-3 region of S. Typhimurium instead of its own SPI-3. We observed that under stressful conditions the hybrid strain showed a clear impairment in resistance to hydrogen peroxide and decreased survival within U937 culture monocytes. We hypothesized that the marT-fidL operon, encoded in SPI-3, was responsible for the new phenotypes because marT is a pseudogen in S. Typhi and has a demonstrated role as a transcriptional regulator in S. Typhimurium. Therefore we cloned and transferred the S. Typhimurium marT-fidL operon into S. Typhi and confirmed that invasion of monocytes was dramatically decreased. Finally, our findings suggest that the genomic and functional differences between SPI-3 sequences have implications in the host specificity of Typhi and Typhimurium serovars.
Conflict of interest statement
Figures



Similar articles
-
Lose to win: marT pseudogenization in Salmonella enterica serovar Typhi contributed to the surV-dependent survival to H2O2, and inside human macrophage-like cells.Infect Genet Evol. 2016 Nov;45:111-121. doi: 10.1016/j.meegid.2016.08.029. Epub 2016 Aug 24. Infect Genet Evol. 2016. PMID: 27567490
-
The Salmonella typhi melittin resistance gene pqaB affects intracellular growth in PMA-differentiated U937 cells, polymyxin B resistance and lipopolysaccharide.Microbiology (Reading). 1999 Feb;145 ( Pt 2):367-378. doi: 10.1099/13500872-145-2-367. Microbiology (Reading). 1999. PMID: 10075419
-
The Salmonella Typhi hlyE gene plays a role in invasion of cultured epithelial cells and its functional transfer to S. Typhimurium promotes deep organ infection in mice.Res Microbiol. 2008 May;159(4):279-87. doi: 10.1016/j.resmic.2008.02.006. Epub 2008 Mar 16. Res Microbiol. 2008. PMID: 18434098
-
Evolutionary strategies of human pathogens.Cold Spring Harb Symp Quant Biol. 2003;68:151-8. doi: 10.1101/sqb.2003.68.151. Cold Spring Harb Symp Quant Biol. 2003. PMID: 15338613 Review. No abstract available.
-
So similar, yet so different: uncovering distinctive features in the genomes of Salmonella enterica serovars Typhimurium and Typhi.FEMS Microbiol Lett. 2010 Apr;305(1):1-13. doi: 10.1111/j.1574-6968.2010.01904.x. Epub 2010 Jan 20. FEMS Microbiol Lett. 2010. PMID: 20146749 Review.
Cited by
-
Helicobacter and salmonella persistent infection strategies.Cold Spring Harb Perspect Med. 2013 Dec 1;3(12):a010348. doi: 10.1101/cshperspect.a010348. Cold Spring Harb Perspect Med. 2013. PMID: 24296347 Free PMC article. Review.
-
Genomic Comparison of the Closely-Related Salmonella enterica Serovars Enteritidis, Dublin and Gallinarum.PLoS One. 2015 Jun 3;10(6):e0126883. doi: 10.1371/journal.pone.0126883. eCollection 2015. PLoS One. 2015. PMID: 26039056 Free PMC article.
-
Comprehensive assignment of roles for Salmonella typhimurium genes in intestinal colonization of food-producing animals.PLoS Genet. 2013 Apr;9(4):e1003456. doi: 10.1371/journal.pgen.1003456. Epub 2013 Apr 18. PLoS Genet. 2013. PMID: 23637626 Free PMC article.
-
Draft Genome Sequence of Salmonella enterica Serovar Typhi Strain STH2370.Genome Announc. 2014 Feb 20;2(1):e00104-14. doi: 10.1128/genomeA.00104-14. Genome Announc. 2014. PMID: 24558245 Free PMC article.
-
Transcriptional regulator MarT negatively regulates MarT-regulated motility gene I, a new gene involved in invasion and virulence of Salmonella enterica.Front Microbiol. 2024 Aug 15;15:1430982. doi: 10.3389/fmicb.2024.1430982. eCollection 2024. Front Microbiol. 2024. PMID: 39211323 Free PMC article.
References
-
- Baker S, Dougan G. The genome of Salmonella enterica serovar Typhi. Clin Infect Dis. 2007;45(Suppl 1):S29–33. - PubMed
-
- Retamal P, Castillo-Ruiz M, Mora GC. Characterization of MgtC, a Virulence Factor of Salmonella enterica Serovar Typhi. PLoS ONE. 2009;4:e5551. doi: 5510.1371/journal.pone.0005551. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

