Analysis of the interactions between the C-terminal cytoplasmic domains of KCNQ1 and KCNE1 channel subunits
- PMID: 20196769
- PMCID: PMC2888147
- DOI: 10.1042/BJ20090977
Analysis of the interactions between the C-terminal cytoplasmic domains of KCNQ1 and KCNE1 channel subunits
Abstract
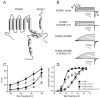
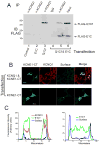
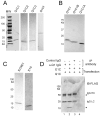
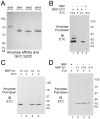
Ion channel subunits encoded by KCNQ1 and KCNE1 produce the slowly activating K+ current (IKs) that plays a central role in myocardial repolarization. The KCNQ1 alpha-subunit and the KCNE1 beta-subunit assemble with their membrane-spanning segments interacting, resulting in transformation of channel activation kinetics. We recently reported a functional interaction involving C-terminal portions of the two subunits with ensuing regulation of channel deactivation. In the present study, we provide evidence characterizing a physical interaction between the KCNQ1-CT (KCNE1 C-terminus) and the KCNE1-CT (KCNE1 C-terminus). When expressed in cultured cells, the KCNE1-CT co-localized with KCNQ1, co-immunoprecipitated with KCNQ1 and perturbed deactivation kinetics of the KCNQ1 currents. Purified KCNQ1-CT and KCNE1-CT physically interacted in pull-down experiments, indicating a direct association. Deletion analysis of KCNQ1-CT indicated that the KCNE1-CT binds to a KCNQ1 region just after the last transmembrane segment, but N-terminal to the tetramerization domain. SPR (surface plasmon resonance) corroborated the pull-down results, showing that the most proximal region (KCNQ1 amino acids 349-438) contributed most to the bimolecular interaction with a dissociation constant of approximately 4 microM. LQT (long QT) mutants of the KCNE1-CT, D76N and W87F, retained binding to the KCNQ1-CT with comparable affinity, indicating that these disease-causing mutations do not alter channel behaviour by disruption of the association. Several LQT mutations involving the KCNQ1-CT, however, showed various effects on KCNQ1/KCNE1 association. Our results indicate that the KCNQ1-CT and the KCNE1-CT comprise an independent interaction domain that may play a role in IKs channel regulation that is potentially affected in some LQTS (LQT syndrome) mutations.
Figures


References
-
- Robbins J. KCNQ potassium channels: physiology, pathophysiology, and pharmacology. Pharmacol Ther. 2001;90:1–19. - PubMed
-
- Barhanin J, Lesage F, Guillemare E, Fink M, Lazdunski M, Romey G. KVLQT1 and IsK (minK) proteins associate to form IKs cardiac potassium current. Nature. 1996;384:78–80. - PubMed
-
- Sanguinetti MC, Curran ME, Zou A, Shen J, Spector PS, Atkinson DL, Keating MT. Coassembly of KVLQT1 and minK (IsK) proteins to form cardiac IKs potassium channel. Nature. 1996;384:80–83. - PubMed
-
- Keating MT, Sanguinetti MC. Molecular and cellular mechanisms of cardiac arrhythmias. Cell. 2001;104:569–580. - PubMed
-
- Splawski I, Tristani-Firouzi M, Lehmann MH, Sanguinetti MC, Keating MT. Mutations in the hminK gene cause long QT syndrome and suppress IKs function. Nat Genet. 1997;17:338–340. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

