Identification and characterization of Dlc1 isoforms in the mouse and study of the biological function of a single gene trapped isoform
- PMID: 20199662
- PMCID: PMC2839985
- DOI: 10.1186/1741-7007-8-17
Identification and characterization of Dlc1 isoforms in the mouse and study of the biological function of a single gene trapped isoform
Abstract
Background: The Dlc1 (deleted in liver cancer 1) tumour suppressor gene codes for a RhoGTPase activating protein that is found inactivated in many tumour types. Several transcriptional isoforms have been described but the functional significance and tissue distribution of each form is presently poorly understood. Also, differences in the number of isoforms and splice variants reported still exist between different mammalian species. In order to better understand the number and function of the different variants of the Dlc1 gene in the mouse, we have carried out a detailed analysis. Extensive 3' RACE experiments were carried out in order to identify all possible Dlc1 isoforms and splice variants in the mouse. In addition, we have generated a gene trapped mouse that targets one of these isoforms in order to study its biological function. The effect of this gene trap insertion on the splicing of other isoforms has also been studied.
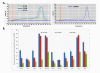
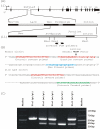
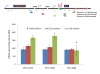
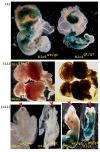
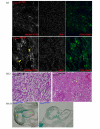
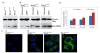
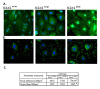
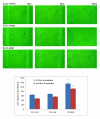
Results: In addition to the known 6.1 and 6.2 Kb transcripts of Dlc1, our study revealed the existence of a novel 7.6 Kb transcriptional isoform in the mouse, which corresponds to the human 7.4 Kb (KIAA1723) cDNA transcript. A gene trapped embryonic cell line, with an insertion between Exon 1 and 2 of the 6.1 Kb transcriptional isoform, was used to generate a transgenic mouse. This line showed a significant reduction in the expression of the trapped isoform. However, reduced expression of the other isoforms was not seen. Mice heterozygous for the gene trapped allele were phenotypically normal, but homozygous mutant embryos did not survive beyond 10.5 days post coitum. Dlc1gt/gt embryos showed defects in the brain, heart, and placental blood vessels. Cultured serum-free mouse embryo cells from Dlc1 deficient embryos had elevated RhoA activity and displayed alterations in the organization of actin filaments and focal adhesions. The Dlc1 deficient cells also exhibited increased wound closure in an in vitro scratch assay.
Conclusions: The mouse has three major transcriptional isoforms of the Dlc1 gene that are differentially expressed in various tissues. A mouse with exon 1 of the 6.1 Kb transcript gt resulted in hypomorphic expression of Dlc1 protein and an embryonic lethal phenotype in the homozygous condition, which indicates that this isoform plays a major role in mouse development. The Dlc1 deficient cells showed altered cytoskeleton structure, increased RhoA activity and cellular migration.
Figures






Similar articles
-
The role of Dlc1 isoform 2 in K-Ras2(G12D) induced thymic cancer.PLoS One. 2012;7(7):e40302. doi: 10.1371/journal.pone.0040302. Epub 2012 Jul 5. PLoS One. 2012. PMID: 22792269 Free PMC article.
-
Aberrant DNA methylation of alternative promoter of DLC1 isoform 1 in meningiomas.J Neurooncol. 2016 Dec;130(3):473-484. doi: 10.1007/s11060-016-2261-3. Epub 2016 Sep 10. J Neurooncol. 2016. PMID: 27614886 Free PMC article.
-
DLC-1, a Rho GTPase-activating protein with tumor suppressor function, is essential for embryonic development.FEBS Lett. 2005 Feb 14;579(5):1191-6. doi: 10.1016/j.febslet.2004.12.090. Epub 2005 Jan 19. FEBS Lett. 2005. PMID: 15710412
-
Tumor suppressor gene DLC1: Its modifications, interactive molecules, and potential prospects for clinical cancer application.Int J Biol Macromol. 2021 Jul 1;182:264-275. doi: 10.1016/j.ijbiomac.2021.04.022. Epub 2021 Apr 6. Int J Biol Macromol. 2021. PMID: 33836193 Review.
-
Deleted in liver cancer-1 (DLC1): an emerging metastasis suppressor gene.Mol Diagn Ther. 2014 Jun;18(3):293-302. doi: 10.1007/s40291-014-0086-3. Mol Diagn Ther. 2014. PMID: 24519699 Free PMC article. Review.
Cited by
-
A discovery resource of rare copy number variations in individuals with autism spectrum disorder.G3 (Bethesda). 2012 Dec;2(12):1665-85. doi: 10.1534/g3.112.004689. Epub 2012 Dec 1. G3 (Bethesda). 2012. PMID: 23275889 Free PMC article.
-
Dlc1 interaction with non-muscle myosin heavy chain II-A (Myh9) and Rac1 activation.Biol Open. 2016 Apr 15;5(4):452-60. doi: 10.1242/bio.015859. Biol Open. 2016. PMID: 26977077 Free PMC article.
-
Inactivation of the Dlc1 gene cooperates with downregulation of p15INK4b and p16Ink4a, leading to neoplastic transformation and poor prognosis in human cancer.Cancer Res. 2012 Nov 15;72(22):5900-11. doi: 10.1158/0008-5472.CAN-12-2368. Epub 2012 Sep 25. Cancer Res. 2012. PMID: 23010077 Free PMC article.
-
CDK5 is a major regulator of the tumor suppressor DLC1.J Cell Biol. 2014 Dec 8;207(5):627-42. doi: 10.1083/jcb.201405105. Epub 2014 Dec 1. J Cell Biol. 2014. PMID: 25452387 Free PMC article.
-
Whole exome sequencing identifies novel predisposing genes in neural tube defects.Mol Genet Genomic Med. 2019 Jan;7(1):e00467. doi: 10.1002/mgg3.467. Epub 2018 Nov 10. Mol Genet Genomic Med. 2019. PMID: 30415495 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

