LRPPRC and SLIRP interact in a ribonucleoprotein complex that regulates posttranscriptional gene expression in mitochondria
- PMID: 20200222
- PMCID: PMC2854090
- DOI: 10.1091/mbc.e10-01-0047
LRPPRC and SLIRP interact in a ribonucleoprotein complex that regulates posttranscriptional gene expression in mitochondria
Abstract
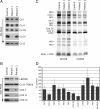
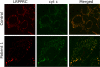
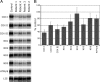
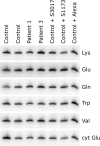
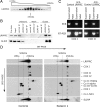
Mutations in LRPPRC are responsible for the French Canadian variant of Leigh syndrome (LSFC), a neurodegenerative disorder caused by a tissue-specific deficiency in cytochrome c oxidase (COX). To investigate the pathogenic mechanism of disease, we studied LRPPRC function in LSFC and control fibroblasts. The level of mutated LRPPRC is reduced in LSFC cells, and this results in decreased steady-state levels of most mitochondrial mRNAs, but not rRNAs or tRNAs, a phenotype that can be reproduced by siRNA-mediated knockdown of LRPPRC in control cells. Processing of the primary transcripts appears normal. The resultant defect in mitochondrial protein synthesis in LSFC cells disproportionately affects the COX subunits, leading to an isolated COX assembly defect. Further knockdown of LRPPRC produces a generalized assembly defect in all oxidative phosphorylation complexes containing mtDNA-encoded subunits, due to a severe decrease in all mitochondrial mRNAs. LRPPRC exists in a high-molecular-weight complex, and it coimmunoprecipitates with SLIRP, a stem-loop RNA-binding protein. Although this interaction does not depend on mitochondrial mRNA, both proteins show reduced stability in its absence. These results implicate LRPPRC in posttranscriptional mitochondrial gene expression as part of a ribonucleoprotein complex that regulates the stability and handling of mature mRNAs.
Figures
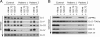


Similar articles
-
Tissue-specific responses to the LRPPRC founder mutation in French Canadian Leigh Syndrome.Hum Mol Genet. 2015 Jan 15;24(2):480-91. doi: 10.1093/hmg/ddu468. Epub 2014 Sep 11. Hum Mol Genet. 2015. PMID: 25214534 Free PMC article.
-
mTORC1 is required for expression of LRPPRC and cytochrome-c oxidase but not HIF-1α in Leigh syndrome French Canadian type patient fibroblasts.Am J Physiol Cell Physiol. 2019 Jul 1;317(1):C58-C67. doi: 10.1152/ajpcell.00160.2017. Epub 2019 Apr 17. Am J Physiol Cell Physiol. 2019. PMID: 30995105 Free PMC article.
-
The role of the LRPPRC (leucine-rich pentatricopeptide repeat cassette) gene in cytochrome oxidase assembly: mutation causes lowered levels of COX (cytochrome c oxidase) I and COX III mRNA.Biochem J. 2004 Aug 15;382(Pt 1):331-6. doi: 10.1042/BJ20040469. Biochem J. 2004. PMID: 15139850 Free PMC article.
-
LRPPRC: A Multifunctional Protein Involved in Energy Metabolism and Human Disease.Front Physiol. 2019 May 24;10:595. doi: 10.3389/fphys.2019.00595. eCollection 2019. Front Physiol. 2019. PMID: 31178748 Free PMC article.
-
Human cytochrome oxidase deficiency.Pediatr Res. 2000 Nov;48(5):581-5. doi: 10.1203/00006450-200011000-00004. Pediatr Res. 2000. PMID: 11044474 Review.
Cited by
-
LRPPRC and SLIRP synergize to maintain sufficient and orderly mammalian mitochondrial translation.Nucleic Acids Res. 2024 Oct 14;52(18):11266-11282. doi: 10.1093/nar/gkae662. Nucleic Acids Res. 2024. PMID: 39087558 Free PMC article.
-
Pathways to balance mitochondrial translation and protein import.Genes Dev. 2018 Oct 1;32(19-20):1285-1296. doi: 10.1101/gad.316547.118. Genes Dev. 2018. PMID: 30275044 Free PMC article. Review.
-
Loss of the nuclear receptor corepressor SLIRP compromises male fertility.PLoS One. 2013 Aug 15;8(8):e70700. doi: 10.1371/journal.pone.0070700. eCollection 2013. PLoS One. 2013. PMID: 23976951 Free PMC article.
-
Elevated levels of mitochondrion-associated autophagy inhibitor LRPPRC are associated with poor prognosis in patients with prostate cancer.Cancer. 2014 Apr 15;120(8):1228-36. doi: 10.1002/cncr.28551. Epub 2014 Jan 3. Cancer. 2014. PMID: 24390809 Free PMC article.
-
Functional genomic analysis of human mitochondrial RNA processing.Cell Rep. 2014 May 8;7(3):918-31. doi: 10.1016/j.celrep.2014.03.035. Epub 2014 Apr 18. Cell Rep. 2014. PMID: 24746820 Free PMC article.
References
-
- Antonicka H., Leary S. C., Guercin G. H., Agar J. N., Horvath R., Kennaway N. G., Harding C. O., Jaksch M., Shoubridge E. A. Mutations in COX10 result in a defect in mitochondrial heme A biosynthesis and account for multiple, early-onset clinical phenotypes associated with isolated COX deficiency. Hum. Mol. Genet. 2003;12:2693–2702. - PubMed
-
- Capaldi R. A., Marusich M. F., Taanman J. W. Mammalian cytochrome-c oxidase: characterization of enzyme and immunological detection of subunits in tissue extracts and whole cells. Methods Enzymol. 1995;260:117–132. - PubMed
-
- Colley S. M., Leedman P. J. SRA and its binding partners: an expanding role for RNA-binding coregulators in nuclear receptor-mediated gene regulation. Crit. Rev. Biochem. Mol. Biol. 2009;44:25–33. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

