Human cytomegalovirus IE2 86 and IE2 40 proteins differentially regulate UL84 protein expression posttranscriptionally in the absence of other viral gene products
- PMID: 20200242
- PMCID: PMC2863830
- DOI: 10.1128/JVI.00090-10
Human cytomegalovirus IE2 86 and IE2 40 proteins differentially regulate UL84 protein expression posttranscriptionally in the absence of other viral gene products
Abstract
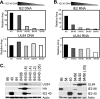
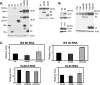
It has previously been demonstrated that, during human cytomegalovirus infection, the viral IE2 86 and IE2 40 proteins are both important for the expression of an early-late viral protein, UL84. Here, we show that expression of the UL84 protein is enhanced upon cotransfection with either IE2 86 or IE2 40, although IE2 40 appears to play a more important role. The UL84 protein levels are tightly linked to the amount of IE2 40 present, but this does not appear to be true for IE2 86. RNA remains constant for all corresponding proteins, indicating posttranscriptional regulation of UL84. The first 105 amino acids of UL84 are necessary and sufficient for this phenotype, and this region is also required for an interaction with IE2 86 and IE2 40. Treatment with proteasome inhibitors shows that UL84 exhibits some proteasome-dependent degradation, and UL84 is not protected against this degradation when coexpressed with IE2 86 or IE2 40. UL84 also exhibits an inhibitory effect on IE2 86 and IE2 40 protein levels in these cotransfection assays. Further, we show that the amino acid sequence of UL84 is important for the enhancement governed by IE2 40. These results indicate that IE2 86, IE2 40, and UL84 serve to regulate protein expression in a posttranscriptional fashion and that this regulation is independent of other viral proteins.
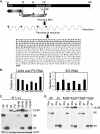
Figures
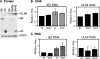



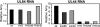
Similar articles
-
Characteristics of Immediate-Early 2 (IE2) and UL84 Proteins in UL84-Independent Strains of Human Cytomegalovirus (HCMV).Microbiol Spectr. 2021 Oct 31;9(2):e0053921. doi: 10.1128/Spectrum.00539-21. Epub 2021 Sep 22. Microbiol Spectr. 2021. PMID: 34550009 Free PMC article.
-
Internal deletions of IE2 86 and loss of the late IE2 60 and IE2 40 proteins encoded by human cytomegalovirus affect the levels of UL84 protein but not the amount of UL84 mRNA or the loading and distribution of the mRNA on polysomes.J Virol. 2008 Nov;82(22):11383-97. doi: 10.1128/JVI.01293-08. Epub 2008 Sep 10. J Virol. 2008. PMID: 18787008 Free PMC article.
-
Human cytomegalovirus DNA replication requires transcriptional activation via an IE2- and UL84-responsive bidirectional promoter element within oriLyt.J Virol. 2004 Nov;78(21):11664-77. doi: 10.1128/JVI.78.21.11664-11677.2004. J Virol. 2004. PMID: 15479808 Free PMC article.
-
Human cytomegalovirus UL84 oligomerization and heterodimerization domains act as transdominant inhibitors of oriLyt-dependent DNA replication: evidence that IE2-UL84 and UL84-UL84 interactions are required for lytic DNA replication.J Virol. 2004 Sep;78(17):9203-14. doi: 10.1128/JVI.78.17.9203-9214.2004. J Virol. 2004. PMID: 15308715 Free PMC article.
-
Nuts and bolts of human cytomegalovirus lytic DNA replication.Curr Top Microbiol Immunol. 2008;325:153-66. doi: 10.1007/978-3-540-77349-8_9. Curr Top Microbiol Immunol. 2008. PMID: 18637505 Review.
Cited by
-
Characteristics of Immediate-Early 2 (IE2) and UL84 Proteins in UL84-Independent Strains of Human Cytomegalovirus (HCMV).Microbiol Spectr. 2021 Oct 31;9(2):e0053921. doi: 10.1128/Spectrum.00539-21. Epub 2021 Sep 22. Microbiol Spectr. 2021. PMID: 34550009 Free PMC article.
-
Identification of compounds with anti-human cytomegalovirus activity that inhibit production of IE2 proteins.Antiviral Res. 2017 Feb;138:61-67. doi: 10.1016/j.antiviral.2016.12.006. Epub 2016 Dec 9. Antiviral Res. 2017. PMID: 27956134 Free PMC article.
-
Mutation of glutamine to arginine at position 548 of IE2 86 in human cytomegalovirus leads to decreased expression of IE2 40, IE2 60, UL83, and UL84 and increased transcription of US8-9 and US29-32.J Virol. 2011 Nov;85(21):11098-110. doi: 10.1128/JVI.05315-11. Epub 2011 Aug 24. J Virol. 2011. PMID: 21865379 Free PMC article.
-
Genome Engineering as a Therapeutic Approach in Cancer Therapy: A Comprehensive Review.Adv Genet (Hoboken). 2024 Feb 5;5(1):2300201. doi: 10.1002/ggn2.202300201. eCollection 2024 Mar. Adv Genet (Hoboken). 2024. PMID: 38465225 Free PMC article. Review.
-
Dynamic and nucleolin-dependent localization of human cytomegalovirus UL84 to the periphery of viral replication compartments and nucleoli.J Virol. 2014 Oct;88(20):11738-47. doi: 10.1128/JVI.01889-14. Epub 2014 Jul 30. J Virol. 2014. PMID: 25078694 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

