Group-specific structural features of the 5'-proximal sequences of coronavirus genomic RNAs
- PMID: 20202661
- PMCID: PMC7111916
- DOI: 10.1016/j.virol.2010.02.007
Group-specific structural features of the 5'-proximal sequences of coronavirus genomic RNAs
Abstract
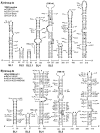
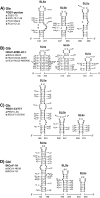
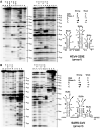
Global predictions of the secondary structure of coronavirus (CoV) 5' untranslated regions and adjacent coding sequences revealed the presence of conserved structural elements. Stem loops (SL) 1, 2, 4, and 5 were predicted in all CoVs, while the core leader transcription-regulating sequence (L-TRS) forms SL3 in only some CoVs. SL5 in group I and II CoVs, with the exception of group IIa CoVs, is characterized by the presence of a large sequence insertion capable of forming hairpins with the conserved 5'-UUYCGU-3' loop sequence. Structure probing confirmed the existence of these hairpins in the group I Human coronavirus-229E and the group II Severe acute respiratory syndrome coronavirus (SARS-CoV). In general, the pattern of the 5' cis-acting elements is highly related to the lineage of CoVs, including features of the conserved hairpins in SL5. The function of these conserved hairpins as a putative packaging signal is discussed.
Figures







References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

