Rare sequence variation in the genome flanking a short tandem repeat locus can lead to a question of "nonmaternity"
- PMID: 20203001
- PMCID: PMC2860477
- DOI: 10.2353/jmoldx.2010.090201
Rare sequence variation in the genome flanking a short tandem repeat locus can lead to a question of "nonmaternity"
Abstract
Typing of STR (short tandem repeat) alleles is used in a variety of applications in clinical molecular pathology, including evaluations for maternal cell contamination. Using a commercially available STR typing assay for maternal cell contamination performed in conjunction with prenatal diagnostic testing, we were posed with apparent nonmaternity when the two fetal samples did not demonstrate the expected maternal allele at one locus. By designing primers external to the region amplified by the primers from the commercial assay and by performing direct sequencing of the resulting amplicon, we were able to determine that a guanine to adenine sequence variation led to primer mismatch and allele dropout. This explained the apparent null allele shared between the maternal and fetal samples. Therefore, although rare, allele dropout must be considered whenever unexplained homozygosity at an STR locus is observed.
Figures
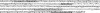

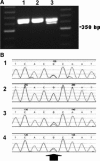

References
-
- Adamson D, Albertson H, Ballard L, Bradley P, Carlson M, Cartwright P, Council C, Elsner T, Fuhrman D, Gerken S, Harris L, Holik P, Kimball A, Knell J, Lawrence E, Lu J, Marks A, Matsunami N, Melis R, Milner B, Moore M, Nelson L, Odelberg R, Peters G, Plaetke R, Riley R, Robertson M, Sargent R, Staker G, Tingey A, Ward K, Zhao X, White R. A collection of ordered tetranucleotide-repeat markers from the human genome. Am J Hum Genet. 1995;57:619–628. - PMC - PubMed
-
- Kimpton CP, Gill P, Walton A, Urquhart A, Millican ES, Adams M. Automated DNA profiling employing multiplex amplification of short tandem repeat loci. PCR Methods Appl. 1993;3:13–22. - PubMed
-
- Urquhart A, Kimpton CP, Downes TJ, Gill P. Variation in short tandem repeat sequences: a survey of twelve microsatellite loci for use as forensic identification markers. Int J Legal Med. 1994;107:13–20. - PubMed
-
- Butler JM. Short tandem repeat typing technologies used in human identity testing. BioTechniques. 2007;43:ii–iv. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

