BglBricks: A flexible standard for biological part assembly
- PMID: 20205762
- PMCID: PMC2822740
- DOI: 10.1186/1754-1611-4-1
BglBricks: A flexible standard for biological part assembly
Abstract
Background: Standard biological parts, such as BioBricks parts, provide the foundation for a new engineering discipline that enables the design and construction of synthetic biological systems with a variety of applications in bioenergy, new materials, therapeutics, and environmental remediation. Although the original BioBricks assembly standard has found widespread use, it has several shortcomings that limit its range of potential applications. In particular, the system is not suitable for the construction of protein fusions due to an unfavorable scar sequence that encodes an in-frame stop codon.
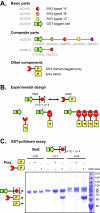
Results: Here, we present a similar but new composition standard, called BglBricks, that addresses the scar translation issue associated with the original standard. The new system employs BglII and BamHI restriction enzymes, robust cutters with an extensive history of use, and results in a 6-nucleotide scar sequence encoding glycine-serine, an innocuous peptide linker in most protein fusion applications. We demonstrate the utility of the new standard in three distinct applications, including the construction of constitutively active gene expression devices with a wide range of expression profiles, the construction of chimeric, multi-domain protein fusions, and the targeted integration of functional DNA sequences into specific loci of the E. coli genome.
Conclusions: The BglBrick standard provides a new, more flexible platform from which to generate standard biological parts and automate DNA assembly. Work on BglBrick assembly reactions, as well as on the development of automation and bioinformatics tools, is currently underway. These tools will provide a foundation from which to transform genetic engineering from a technically intensive art into a purely design-based discipline.
Figures



References
-
- Knight TF. , JrIdempotent Vector Design for Standard Assembly of Biobricks. DSpace. 2003. http://hdl.handle.net/1721.1/21168
-
- Registry of Standard Biological Parts. http://www.partsregistry.org
-
- Brown J. The iGEM competition: building with biology. IET Synth Biol. 2007;1:3–6. doi: 10.1049/iet-stb:20079020. - DOI
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

