Cell type-specific gene expression differences in complex tissues
- PMID: 20208531
- PMCID: PMC3699332
- DOI: 10.1038/nmeth.1439
Cell type-specific gene expression differences in complex tissues
Abstract
We describe cell type-specific significance analysis of microarrays (csSAM) for analyzing differential gene expression for each cell type in a biological sample from microarray data and relative cell-type frequencies. First, we validated csSAM with predesigned mixtures and then applied it to whole-blood gene expression datasets from stable post-transplant kidney transplant recipients and those experiencing acute transplant rejection, which revealed hundreds of differentially expressed genes that were otherwise undetectable.
Figures
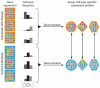

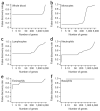
Comment in
-
Gene expression deconvolution in linear space.Nat Methods. 2011 Dec 28;9(1):8-9; author reply 9. doi: 10.1038/nmeth.1830. Nat Methods. 2011. PMID: 22205510 No abstract available.
References
Publication types
MeSH terms
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

