Differentiation of symbiotic cells and endosymbionts in Medicago truncatula nodulation are coupled to two transcriptome-switches
- PMID: 20209049
- PMCID: PMC2832008
- DOI: 10.1371/journal.pone.0009519
Differentiation of symbiotic cells and endosymbionts in Medicago truncatula nodulation are coupled to two transcriptome-switches
Abstract
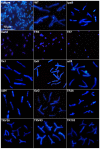
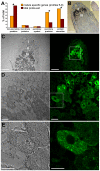
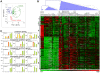
The legume plant Medicago truncatula establishes a symbiosis with the nitrogen-fixing bacterium Sinorhizobium meliloti which takes place in root nodules. The formation of nodules employs a complex developmental program involving organogenesis, specific cellular differentiation of the host cells and the endosymbiotic bacteria, called bacteroids, as well as the specific activation of a large number of plant genes. By using a collection of plant and bacterial mutants inducing non-functional, Fix(-) nodules, we studied the differentiation processes of the symbiotic partners together with the nodule transcriptome, with the aim of unravelling links between cell differentiation and transcriptome activation. Two waves of transcriptional reprogramming involving the repression and the massive induction of hundreds of genes were observed during wild-type nodule formation. The dominant features of this "nodule-specific transcriptome" were the repression of plant defense-related genes, the transient activation of cell cycle and protein synthesis genes at the early stage of nodule development and the activation of the secretory pathway along with a large number of transmembrane and secretory proteins or peptides throughout organogenesis. The fifteen plant and bacterial mutants that were analyzed fell into four major categories. Members of the first category of mutants formed non-functional nodules although they had differentiated nodule cells and bacteroids. This group passed the two transcriptome switch-points similarly to the wild type. The second category, which formed nodules in which the plant cells were differentiated and infected but the bacteroids did not differentiate, passed the first transcriptome switch but not the second one. Nodules in the third category contained infection threads but were devoid of differentiated symbiotic cells and displayed a root-like transcriptome. Nodules in the fourth category were free of bacteria, devoid of differentiated symbiotic cells and also displayed a root-like transcriptome. A correlation thus exists between the differentiation of symbiotic nodule cells and the first wave of nodule specific gene activation and between differentiation of rhizobia to bacteroids and the second transcriptome wave in nodules. The differentiation of symbiotic cells and of bacteroids may therefore constitute signals for the execution of these transcriptome-switches.
Conflict of interest statement
Figures




Similar articles
-
Transcriptomic Analysis of Sinorhizobium meliloti and Medicago truncatula Symbiosis Using Nitrogen Fixation-Deficient Nodules.Mol Plant Microbe Interact. 2015 Aug;28(8):856-68. doi: 10.1094/MPMI-12-14-0407-R. Epub 2015 Jul 16. Mol Plant Microbe Interact. 2015. PMID: 25844838
-
Sinorhizobium meliloti Functions Required for Resistance to Antimicrobial NCR Peptides and Bacteroid Differentiation.mBio. 2021 Aug 31;12(4):e0089521. doi: 10.1128/mBio.00895-21. Epub 2021 Jul 27. mBio. 2021. PMID: 34311575 Free PMC article.
-
From Intracellular Bacteria to Differentiated Bacteroids: Transcriptome and Metabolome Analysis in Aeschynomene Nodules Using the Bradyrhizobium sp. Strain ORS285 bclA Mutant.J Bacteriol. 2019 Aug 8;201(17):e00191-19. doi: 10.1128/JB.00191-19. Print 2019 Sep 1. J Bacteriol. 2019. PMID: 31182497 Free PMC article.
-
Gene Expression in Nitrogen-Fixing Symbiotic Nodule Cells in Medicago truncatula and Other Nodulating Plants.Plant Cell. 2020 Jan;32(1):42-68. doi: 10.1105/tpc.19.00494. Epub 2019 Nov 11. Plant Cell. 2020. PMID: 31712407 Free PMC article. Review.
-
Exploring the role of symbiotic modifier peptidases in the legume - rhizobium symbiosis.Arch Microbiol. 2024 Mar 11;206(4):147. doi: 10.1007/s00203-024-03920-w. Arch Microbiol. 2024. PMID: 38462552 Review.
Cited by
-
Failure of self-control: defense-like reactions during legume/rhizobia symbiosis.Plant Signal Behav. 2013 Apr;8(4):e23915. doi: 10.4161/psb.23915. Epub 2013 Feb 20. Plant Signal Behav. 2013. PMID: 23425859 Free PMC article.
-
Regulation of Small RNAs and Corresponding Targets in Nod Factor-Induced Phaseolus vulgaris Root Hair Cells.Int J Mol Sci. 2016 Jun 4;17(6):887. doi: 10.3390/ijms17060887. Int J Mol Sci. 2016. PMID: 27271618 Free PMC article.
-
NODULE ROOT and COCHLEATA maintain nodule development and are legume orthologs of Arabidopsis BLADE-ON-PETIOLE genes.Plant Cell. 2012 Nov;24(11):4498-510. doi: 10.1105/tpc.112.103747. Epub 2012 Nov 6. Plant Cell. 2012. PMID: 23136374 Free PMC article.
-
Next-generation annotation of prokaryotic genomes with EuGene-P: application to Sinorhizobium meliloti 2011.DNA Res. 2013 Aug;20(4):339-54. doi: 10.1093/dnares/dst014. Epub 2013 Apr 18. DNA Res. 2013. PMID: 23599422 Free PMC article.
-
In-planta Sporulation Capacity Enhances Infectivity and Rhizospheric Competitiveness of Frankia Strains.Microbes Environ. 2016;31(1):11-8. doi: 10.1264/jsme2.ME15090. Epub 2015 Dec 26. Microbes Environ. 2016. PMID: 26726131 Free PMC article.
References
-
- Maunoury N, Kondorosi A, Kondorosi E, Mergaert P. James EK, Sprent JI, Dilworth WE, Newton WE, editors. Cell biology of nodule infection and development. 2008. pp. 153–189. Nitrogen-fixing Leguminous Symbioses: Springer.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

