Identification of two distinct cell binding sequences in the vitamin D binding protein
- PMID: 20211661
- PMCID: PMC2856814
- DOI: 10.1016/j.bbamcr.2010.02.010
Identification of two distinct cell binding sequences in the vitamin D binding protein
Abstract
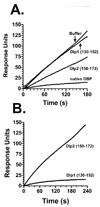
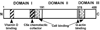
The vitamin D binding protein (DBP) is a multifunctional, albumin-like plasma protein that often requires cell surface binding to mediate some of its diverse functions. DBP binds to several different molecules on the external face of the plasma membrane indicating that it may possess distinct cell binding sequences. In this report, surface plasmon resonance was utilized to evaluate the relative binding of the human myeloid cell line U937 to immobilized recombinant expressed DBP in order to identify cell localization sequences. U937 cells showed robust binding to immobilized native DBP, but essentially no interaction when sensor chips were coated with beta(2)-microglobulin or BSA. The cell-DBP interaction was completely eliminated if cells were pretreated with soluble DBP. Recombinant DBP domains and truncated domains were next evaluated to determine the location of cell binding regions. Domains I (amino acids 1-191) and III (379-458), but not domain II (192-378), could support cell binding. Further evaluation of domain I, using truncated proteins and overlapping peptides, demonstrated that a single amino acid sequence, residues 150-172 (NYGQAPLSLLVSYTKSYLSMVGS), mediated cell binding. The domain III cell binding region was investigated using truncated versions of domain III fused to full-length domain II that served as a scaffold. These experiments indicated that the cell binding sequence is located in the first portion of that domain (379-402: ELSSFIDKGQELCADYSENTFTEY). Overlapping peptides spanning this sequence could partially block cell binding only when used in combination. We conclude that DBP contains two cell localization sequences that may be required for some of the multiple functions of this protein.
Copyright 2010 Elsevier B.V. All rights reserved.
Figures





Similar articles
-
Identification of a region in the vitamin D-binding protein that mediates its C5a chemotactic cofactor function.J Biol Chem. 2004 Dec 17;279(51):53282-7. doi: 10.1074/jbc.M411462200. Epub 2004 Oct 14. J Biol Chem. 2004. PMID: 15485893
-
Direct binding of a fragment of the Wiskott-Aldrich syndrome protein to the C-terminal end of the anaphylatoxin C5a receptor.Biochem J. 2003 Jun 1;372(Pt 2):453-63. doi: 10.1042/BJ20021803. Biochem J. 2003. PMID: 12600272 Free PMC article.
-
CD44 and annexin A2 mediate the C5a chemotactic cofactor function of the vitamin D binding protein.J Immunol. 2005 Oct 1;175(7):4754-60. doi: 10.4049/jimmunol.175.7.4754. J Immunol. 2005. PMID: 16177123
-
Group-specific component is not only a vitamin-D-binding protein.Exp Clin Immunogenet. 1992;9(3):161-75. Exp Clin Immunogenet. 1992. PMID: 1303095 Review.
-
Vitamin D binding protein: genomic structure, functional domains, and mRNA expression in tissues.J Steroid Biochem Mol Biol. 1991;40(4-6):787-93. doi: 10.1016/0960-0760(91)90304-n. J Steroid Biochem Mol Biol. 1991. PMID: 1958576 Review.
Cited by
-
Vitamin D binding protein in psychiatric and neurological disorders: Implications for diagnosis and treatment.Genes Dis. 2024 Apr 15;11(5):101309. doi: 10.1016/j.gendis.2024.101309. eCollection 2024 Sep. Genes Dis. 2024. PMID: 38983447 Free PMC article. Review.
-
Vitamin D binding protein isoforms as candidate predictors of disease extension in childhood arthritis.J Proteomics. 2012 Sep 18;75(17):5479-92. doi: 10.1016/j.jprot.2012.06.024. Epub 2012 Jul 6. J Proteomics. 2012. PMID: 22771520 Free PMC article.
-
Vitamin D and COVID-19: An Overview of Recent Evidence.Int J Mol Sci. 2021 Sep 29;22(19):10559. doi: 10.3390/ijms221910559. Int J Mol Sci. 2021. PMID: 34638897 Free PMC article. Review.
-
Neutrophil recruitment to the lung in both C5a- and CXCL1-induced alveolitis is impaired in vitamin D-binding protein-deficient mice.J Immunol. 2013 Jul 15;191(2):848-56. doi: 10.4049/jimmunol.1202941. Epub 2013 Jun 10. J Immunol. 2013. PMID: 23752613 Free PMC article.
-
Common variants of the vitamin D binding protein gene and adverse health outcomes.Crit Rev Clin Lab Sci. 2013 Jan-Feb;50(1):1-22. doi: 10.3109/10408363.2012.750262. Epub 2013 Feb 22. Crit Rev Clin Lab Sci. 2013. PMID: 23427793 Free PMC article. Review.
References
-
- White P, Cooke N. The multifunctional properties and characteristics of vitamin D-binding protein. Trends Endocrinol Metab. 2000;11:320–327. - PubMed
-
- Gomme PT, Bertolini J. Therapeutic potential of vitamin D-binding protein. Trends Biotechnol. 2004;22:340–345. - PubMed
-
- Cooke NE, Haddad JG. Vitamin D binding protein (Gc-globulin) Endocr.Rev. 1989;10:294–307. - PubMed
-
- Haddad JG, Hu YZ, Kowalski MA, Laramore C, Ray K, Robzyk P, Cooke NE. Identification of the sterol- and actin-binding domains of plasma vitamin D binding protein (Gc-globulin) Biochemistry. 1992;31:7174–7181. - PubMed
-
- Swamy N, Dutta A, Ray R. Roles Of the Structure and Orientation Of Ligands and Ligand Mimics Inside the Ligand-Binding Pocket Of the Vitamin D-Binding Protein. Biochemistry. 1997;36:7432–7436. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

