Identification of hypermethylated genes associated with cisplatin resistance in human cancers
- PMID: 20215521
- PMCID: PMC2849007
- DOI: 10.1158/0008-5472.CAN-09-3427
Identification of hypermethylated genes associated with cisplatin resistance in human cancers
Abstract
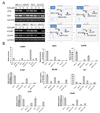
Cisplatin is among the most widely used cytotoxic anticancer agents in solid tumors; however, the development of secondary resistance remains a major obstacle to clinical efficacy. Treatment-related DNA hypermethylation may play a role in creating drug-resistant phenotypes by inactivating genes that are required for cytotoxicity. We applied a pharmacologic unmasking approach to detect hypermethylated genes whose inactivation contributes to cisplatin resistance. Using three pairs of isogeneic, cisplatin-sensitive, and cisplatin-resistant cell lines derived from two parental cell lines (KB-3-1 and SCC25), we identified several hundred genes that were downregulated in each resistant cell line and reactivated by the DNA methyltransferase inhibitor 5-aza-2'-deoxycytidine. Among them, 30 genes were common to two or more cell lines and/or reported to be downregulated in previous studies. Bisulfite sequencing confirmed that 14 genes were hypermethylated in resistant cell lines but not in the sensitive parental cell lines. Six of 14 genes (SAT, C8orf4, LAMB3, TUBB, G0S2, and MCAM) were cisplatin inducible in sensitive but not in resistant cell lines. Small interfering RNA knockdown of two genes, SAT and S100P, increased cell viability with cisplatin treatment in sensitive parental cell lines. S100P knockdown significantly decreased the S-phase fraction of parental sensitive cell lines and slowed cell proliferation, which was associated with decreased sensitivity to cisplatin. Based on these findings, we conclude that DNA methylation is a frequent event in cells that are chronically exposed to cisplatin and that methylation-induced gene silencing may play a role in the development of resistance to cytotoxic chemotherapeutic agents.
Conflict of interest statement
Figures




References
-
- Kruh GD. Introduction to resistance to anticancer agents. Oncogene. 2003;22:7262–7264. - PubMed
-
- Giaccone G. Clinical perspectives on platinum resistance. Drugs. 2000;59(Suppl 4):9–17. discussion 37-8. - PubMed
-
- Dammann R, Li C, Yoon JH, et al. Epigenetic inactivation of a RAS association domain family protein from the lung tumour suppressor locus 3p21.3. Nat Genet. 2000;25:315–319. - PubMed
-
- Li QL, Ito K, Sakakura C, et al. Causal relationship between the loss of RUNX3 expression and gastric cancer. Cell. 2002;109:113–124. - PubMed
-
- Yoshikawa H, Matsubara K, Qian GS, et al. SOCS-1, a negative regulator of the JAK/STAT pathway, is silenced by methylation in human hepatocellular carcinoma and shows growth-suppression activity. Nat Genet. 2001;28:29–35. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

