Proteomic analysis of tardigrades: towards a better understanding of molecular mechanisms by anhydrobiotic organisms
- PMID: 20224743
- PMCID: PMC2835947
- DOI: 10.1371/journal.pone.0009502
Proteomic analysis of tardigrades: towards a better understanding of molecular mechanisms by anhydrobiotic organisms
Abstract
Background: Tardigrades are small, multicellular invertebrates which are able to survive times of unfavourable environmental conditions using their well-known capability to undergo cryptobiosis at any stage of their life cycle. Milnesium tardigradum has become a powerful model system for the analysis of cryptobiosis. While some genetic information is already available for Milnesium tardigradum the proteome is still to be discovered.
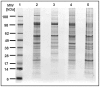
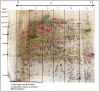
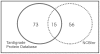
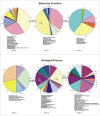
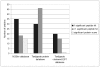
Principal findings: Here we present to the best of our knowledge the first comprehensive study of Milnesium tardigradum on the protein level. To establish a proteome reference map we developed optimized protocols for protein extraction from tardigrades in the active state and for separation of proteins by high resolution two-dimensional gel electrophoresis. Since only limited sequence information of M. tardigradum on the genome and gene expression level is available to date in public databases we initiated in parallel a tardigrade EST sequencing project to allow for protein identification by electrospray ionization tandem mass spectrometry. 271 out of 606 analyzed protein spots could be identified by searching against the publicly available NCBInr database as well as our newly established tardigrade protein database corresponding to 144 unique proteins. Another 150 spots could be identified in the tardigrade clustered EST database corresponding to 36 unique contigs and ESTs. Proteins with annotated function were further categorized in more detail by their molecular function, biological process and cellular component. For the proteins of unknown function more information could be obtained by performing a protein domain annotation analysis. Our results include proteins like protein member of different heat shock protein families and LEA group 3, which might play important roles in surviving extreme conditions.
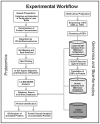
Conclusions: The proteome reference map of Milnesium tardigradum provides the basis for further studies in order to identify and characterize the biochemical mechanisms of tolerance to extreme desiccation. The optimized proteomics workflow will enable application of sensitive quantification techniques to detect differences in protein expression, which are characteristic of the active and anhydrobiotic states of tardigrades.
Conflict of interest statement
Figures




Similar articles
-
Comparative proteome analysis of Milnesium tardigradum in early embryonic state versus adults in active and anhydrobiotic state.PLoS One. 2012;7(9):e45682. doi: 10.1371/journal.pone.0045682. Epub 2012 Sep 27. PLoS One. 2012. PMID: 23029181 Free PMC article.
-
Towards decrypting cryptobiosis--analyzing anhydrobiosis in the tardigrade Milnesium tardigradum using transcriptome sequencing.PLoS One. 2014 Mar 20;9(3):e92663. doi: 10.1371/journal.pone.0092663. eCollection 2014. PLoS One. 2014. PMID: 24651535 Free PMC article.
-
Transcriptome survey of the anhydrobiotic tardigrade Milnesium tardigradum in comparison with Hypsibius dujardini and Richtersius coronifer.BMC Genomics. 2010 Mar 12;11:168. doi: 10.1186/1471-2164-11-168. BMC Genomics. 2010. PMID: 20226016 Free PMC article.
-
Tardigrade stowaways: Literature review of Propyxidium tardigradum (Ciliophora, Peritrichia) and its first record in Scotland.Eur J Protistol. 2023 Jun;89:125974. doi: 10.1016/j.ejop.2023.125974. Epub 2023 Mar 22. Eur J Protistol. 2023. PMID: 37084697 Review.
-
Cryptobiosis: a new theoretical perspective.Prog Biophys Mol Biol. 2006 Oct;92(2):258-67. doi: 10.1016/j.pbiomolbio.2005.11.001. Epub 2005 Dec 9. Prog Biophys Mol Biol. 2006. PMID: 16380155 Review.
Cited by
-
Helicity of a tardigrade disordered protein contributes to its protective function during desiccation.Protein Sci. 2024 Feb;33(2):e4872. doi: 10.1002/pro.4872. Protein Sci. 2024. PMID: 38114424 Free PMC article.
-
Ionizing radiation responses appear incidental to desiccation responses in the bdelloid rotifer Adineta vaga.BMC Biol. 2024 Jan 25;22(1):11. doi: 10.1186/s12915-023-01807-8. BMC Biol. 2024. PMID: 38273318 Free PMC article.
-
Comparative proteome analysis of Milnesium tardigradum in early embryonic state versus adults in active and anhydrobiotic state.PLoS One. 2012;7(9):e45682. doi: 10.1371/journal.pone.0045682. Epub 2012 Sep 27. PLoS One. 2012. PMID: 23029181 Free PMC article.
-
Towards decrypting cryptobiosis--analyzing anhydrobiosis in the tardigrade Milnesium tardigradum using transcriptome sequencing.PLoS One. 2014 Mar 20;9(3):e92663. doi: 10.1371/journal.pone.0092663. eCollection 2014. PLoS One. 2014. PMID: 24651535 Free PMC article.
-
The Physical, Chemical and Physiological Limits of Life.Life (Basel). 2015 Jul 17;5(3):1472-86. doi: 10.3390/life5031472. Life (Basel). 2015. PMID: 26193325 Free PMC article.
References
-
- Keilin D. The problem of anabiosis or latent life: history and current concept. Proc R Soc Lond B Biol Sci. 1959;150:149–191. - PubMed
-
- Baumann H. Die Anabiose der Tardigraden. Zoologischen Jahrbücher. 1922:501–556.
-
- Nelson DR. Current status of the tardigrada: evolution and ecology. Integrative and Comparative Biology. 2002;42:652–659. - PubMed
-
- Schill RO, Fritz GB. Desiccation tolerance in embryonic stages of the tardigrade Minesium tardigradum. Journal of Zoology (London) 2008;276:103–107.
-
- Hengherr S, Brümmer F, Schill RO. Anhydrobiosis in tardigrades and its effects on longevity traits. Journal of Zoology (London): in press 2008
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

