Pathogenic polyglutamine tracts are potent inducers of spontaneous Sup35 and Rnq1 amyloidogenesis
- PMID: 20224794
- PMCID: PMC2835767
- DOI: 10.1371/journal.pone.0009642
Pathogenic polyglutamine tracts are potent inducers of spontaneous Sup35 and Rnq1 amyloidogenesis
Abstract
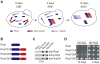
The glutamine/asparagine (Q/N)-rich yeast prion protein Sup35 has a low intrinsic propensity to spontaneously self-assemble into ordered, beta-sheet-rich amyloid fibrils. In yeast cells, de novo formation of Sup35 aggregates is greatly facilitated by high protein concentrations and the presence of preformed Q/N-rich protein aggregates that template Sup35 polymerization. Here, we have investigated whether aggregation-promoting polyglutamine (polyQ) tracts can stimulate the de novo formation of ordered Sup35 protein aggregates in the absence of Q/N-rich yeast prions. Fusion proteins with polyQ tracts of different lengths were produced and their ability to spontaneously self-assemble into amlyloid structures was analyzed using in vitro and in vivo model systems. We found that Sup35 fusions with pathogenic (>or=54 glutamines), as opposed to non-pathogenic (19 glutamines) polyQ tracts efficiently form seeding-competent protein aggregates. Strikingly, polyQ-mediated de novo assembly of Sup35 protein aggregates in yeast cells was independent of pre-existing Q/N-rich protein aggregates. This indicates that increasing the content of aggregation-promoting sequences enhances the tendency of Sup35 to spontaneously self-assemble into insoluble protein aggregates. A similar result was obtained when pathogenic polyQ tracts were linked to the yeast prion protein Rnq1, demonstrating that polyQ sequences are generic inducers of amyloidogenesis. In conclusion, long polyQ sequences are powerful molecular tools that allow the efficient production of seeding-competent amyloid structures.
Conflict of interest statement
Figures



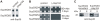


Similar articles
-
Effects of Q/N-rich, polyQ, and non-polyQ amyloids on the de novo formation of the [PSI+] prion in yeast and aggregation of Sup35 in vitro.Proc Natl Acad Sci U S A. 2004 Aug 31;101(35):12934-9. doi: 10.1073/pnas.0404968101. Epub 2004 Aug 23. Proc Natl Acad Sci U S A. 2004. PMID: 15326312 Free PMC article.
-
Heterologous aggregates promote de novo prion appearance via more than one mechanism.PLoS Genet. 2015 Jan 8;11(1):e1004814. doi: 10.1371/journal.pgen.1004814. eCollection 2015 Jan. PLoS Genet. 2015. PMID: 25568955 Free PMC article.
-
Increased [PSI+] appearance by fusion of Rnq1 with the prion domain of Sup35 in Saccharomyces cerevisiae.Eukaryot Cell. 2009 Jul;8(7):968-76. doi: 10.1128/EC.00353-08. Epub 2009 May 1. Eukaryot Cell. 2009. PMID: 19411620 Free PMC article.
-
Prion and nonprion amyloids: a comparison inspired by the yeast Sup35 protein.Prion. 2007 Jul-Sep;1(3):179-84. doi: 10.4161/pri.1.3.4840. Epub 2007 Jul 6. Prion. 2007. PMID: 19164899 Free PMC article. Review.
-
Assembly of the asparagine- and glutamine-rich yeast prions into protein fibrils.Curr Alzheimer Res. 2008 Jun;5(3):251-9. doi: 10.2174/156720508784533303. Curr Alzheimer Res. 2008. PMID: 18537542 Review.
Cited by
-
Location trumps length: polyglutamine-mediated changes in folding and aggregation of a host protein.Biophys J. 2011 Jun 8;100(11):2773-82. doi: 10.1016/j.bpj.2011.04.028. Biophys J. 2011. PMID: 21641323 Free PMC article.
-
[PRION+] States Are Associated with Specific Histone H3 Post-Translational Modification Changes.Pathogens. 2022 Nov 29;11(12):1436. doi: 10.3390/pathogens11121436. Pathogens. 2022. PMID: 36558770 Free PMC article.
-
The Evidence for the Spread and Seeding Capacities of the Mutant Huntingtin Protein in in Vitro Systems and Their Therapeutic Implications.Front Neurosci. 2017 Nov 28;11:647. doi: 10.3389/fnins.2017.00647. eCollection 2017. Front Neurosci. 2017. PMID: 29234268 Free PMC article. Review.
-
Regulated protein aggregation: stress granules and neurodegeneration.Mol Neurodegener. 2012 Nov 20;7:56. doi: 10.1186/1750-1326-7-56. Mol Neurodegener. 2012. PMID: 23164372 Free PMC article. Review.
-
Prion-Like Characteristics of Polyglutamine-Containing Proteins.Cold Spring Harb Perspect Med. 2018 Feb 1;8(2):a024257. doi: 10.1101/cshperspect.a024257. Cold Spring Harb Perspect Med. 2018. PMID: 28096245 Free PMC article. Review.
References
-
- Dobson CM. Protein folding and misfolding. Nature. 2003;426:884–890. - PubMed
-
- Taylor JP, Hardy J, Fischbeck KH. Toxic proteins in neurodegenerative disease. Science. 2002;296:1991–1995. - PubMed
-
- Soto C, Estrada L, Castilla J. Amyloids, prions and the inherent infectious nature of misfolded protein aggregates. Trends Biochem Sci. 2006;31:150–155. - PubMed
-
- Fowler DM, Koulov AV, Balch WE, Kelly JW. Functional amyloid–from bacteria to humans. Trends Biochem Sci. 2007;32:217–224. - PubMed
-
- Bitan G, Lomakin A, Teplow DB. Amyloid beta-protein oligomerization: prenucleation interactions revealed by photo-induced cross-linking of unmodified proteins. J Biol Chem. 2001;276:35176–35184. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

