Characterizing regulatory path motifs in integrated networks using perturbational data
- PMID: 20230615
- PMCID: PMC2864572
- DOI: 10.1186/gb-2010-11-3-r32
Characterizing regulatory path motifs in integrated networks using perturbational data
Abstract
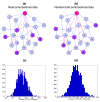
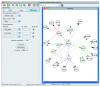
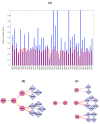
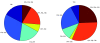
We introduce Pathicular http://bioinformatics.psb.ugent.be/software/details/Pathicular, a Cytoscape plugin for studying the cellular response to perturbations of transcription factors by integrating perturbational expression data with transcriptional, protein-protein and phosphorylation networks. Pathicular searches for 'regulatory path motifs', short paths in the integrated physical networks which occur significantly more often than expected between transcription factors and their targets in the perturbational data. A case study in Saccharomyces cerevisiae identifies eight regulatory path motifs and demonstrates their biological significance.
Figures
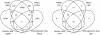

References
-
- Ideker T, Ozier O, Schwikowski B, Siegel AF. Discovering regulatory and signalling circuits in molecular interaction networks. Bioinformatics. 2002;18(Suppl 1):S233–240. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

