Statistical evaluation of transcriptomic data generated using the Affymetrix one-cycle, two-cycle and IVT-Express RNA labelling protocols with the Arabidopsis ATH1 microarray
- PMID: 20230623
- PMCID: PMC2847557
- DOI: 10.1186/1746-4811-6-9
Statistical evaluation of transcriptomic data generated using the Affymetrix one-cycle, two-cycle and IVT-Express RNA labelling protocols with the Arabidopsis ATH1 microarray
Abstract
Background: Microarrays are a powerful tool used for the determination of global RNA expression. There is an increasing requirement to focus on profiling gene expression in tissues where it is difficult to obtain large quantities of material, for example individual tissues within organs such as the root, or individual isolated cells. From such samples, it is difficult to produce the amount of RNA required for labelling and hybridisation in microarray experiments, thus a process of amplification is usually adopted. Despite the increasing use of two-cycle amplification for transcriptomic analyses on the Affymetrix ATH1 array, there has been no report investigating any potential bias in gene representation that may occur as a result.
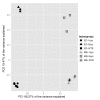
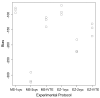
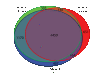
Results: Here we compare transcriptomic data generated using Affymetrix one-cycle (standard labelling protocol), two-cycle (small-sample protocol) and IVT-Express protocols with the Affymetrix ATH1 array using Arabidopsis root samples. Results obtained with each protocol are broadly similar. However, we show that there are 35 probe sets (of a total of 22810) that are misrepresented in the two-cycle data sets. Of these, 33 probe sets were classed as mis-amplified when comparisons of two independent publicly available data sets were undertaken.
Conclusions: Given the unreliable nature of the highlighted probes, we caution against using data associated with the corresponding genes in analyses involving transcriptomic data generated with two-cycle amplification protocols. We have shown that the Affymetrix IVT-E labelling protocol produces data with less associated bias than the two-cycle protocol, and as such, would recommend this kit for new experiments that involve small samples.
Figures
Similar articles
-
Arabidopsis transcript profiling on Affymetrix GeneChip arrays.Plant Mol Biol. 2003 Nov;53(4):457-65. doi: 10.1023/B:PLAN.0000019069.23317.97. Plant Mol Biol. 2003. PMID: 15010612
-
AGRONOMICS1: a new resource for Arabidopsis transcriptome profiling.Plant Physiol. 2010 Feb;152(2):487-99. doi: 10.1104/pp.109.150185. Epub 2009 Dec 23. Plant Physiol. 2010. PMID: 20032078 Free PMC article.
-
An alternative method to amplify RNA without loss of signal conservation for expression analysis with a proteinase DNA microarray in the ArrayTube format.BMC Genomics. 2006 Jun 12;7:144. doi: 10.1186/1471-2164-7-144. BMC Genomics. 2006. PMID: 16768788 Free PMC article.
-
Adjustment method for microarray data generated using two-cycle RNA labeling protocol.BMC Genomics. 2013 Jan 16;14:31. doi: 10.1186/1471-2164-14-31. BMC Genomics. 2013. PMID: 23324182 Free PMC article.
-
Folic acid supplementation and malaria susceptibility and severity among people taking antifolate antimalarial drugs in endemic areas.Cochrane Database Syst Rev. 2022 Feb 1;2(2022):CD014217. doi: 10.1002/14651858.CD014217. Cochrane Database Syst Rev. 2022. PMID: 36321557 Free PMC article.
Cited by
-
AIK1, A Mitogen-Activated Protein Kinase, Modulates Abscisic Acid Responses through the MKK5-MPK6 Kinase Cascade.Plant Physiol. 2017 Feb;173(2):1391-1408. doi: 10.1104/pp.16.01386. Epub 2016 Dec 2. Plant Physiol. 2017. PMID: 27913741 Free PMC article.
-
RootNav: navigating images of complex root architectures.Plant Physiol. 2013 Aug;162(4):1802-14. doi: 10.1104/pp.113.221531. Epub 2013 Jun 13. Plant Physiol. 2013. PMID: 23766367 Free PMC article.
-
Systems analysis of auxin transport in the Arabidopsis root apex.Plant Cell. 2014 Mar;26(3):862-75. doi: 10.1105/tpc.113.119495. Epub 2014 Mar 14. Plant Cell. 2014. PMID: 24632533 Free PMC article.
-
Recovering the dynamics of root growth and development using novel image acquisition and analysis methods.Philos Trans R Soc Lond B Biol Sci. 2012 Jun 5;367(1595):1517-24. doi: 10.1098/rstb.2011.0291. Philos Trans R Soc Lond B Biol Sci. 2012. PMID: 22527394 Free PMC article.
-
CellSeT: novel software to extract and analyze structured networks of plant cells from confocal images.Plant Cell. 2012 Apr;24(4):1353-61. doi: 10.1105/tpc.112.096289. Epub 2012 Apr 3. Plant Cell. 2012. PMID: 22474181 Free PMC article.
References
Grants and funding
LinkOut - more resources
Full Text Sources

