Requirement of the mTOR kinase for the regulation of Maf1 phosphorylation and control of RNA polymerase III-dependent transcription in cancer cells
- PMID: 20233713
- PMCID: PMC2865278
- DOI: 10.1074/jbc.M109.071639
Requirement of the mTOR kinase for the regulation of Maf1 phosphorylation and control of RNA polymerase III-dependent transcription in cancer cells
Abstract
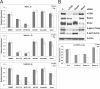
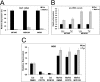
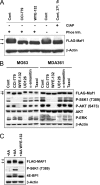
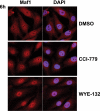
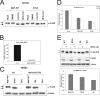
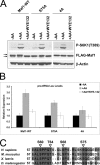
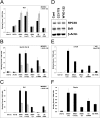
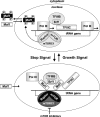
The mammalian target of rapamycin (mTOR) regulates growth via promoting translation and transcription. Here, employing an mTOR active-site inhibitor WYE-125132 (WYE-132), we have performed quantitative phospho-proteomics and identified a Ser-75-containing phosphopeptide from Maf1, a known repressor of RNA polymerase III (Pol III) transcription. Treatment of cancer cells with WYE-132 or the rapamycin analog CCI-779 led to a rapid loss of the phosphorylation at Ser-75, whereas this effect was not seen in cells treated with cytotoxic agents or unrelated inhibitors. WYE-132-induced Maf1 dephosphorylation correlated with its accumulation in the nucleus and a marked decline in the cellular levels of pre-tRNAs. Depletion of cellular Maf1 via small interfering RNA increased basal pre-tRNA and rendered tRNA synthesis refractory to mTOR inhibitors. Maf1 mutant proteins carrying S75A alone or with S60A, T64A, and S68A (Maf1-S75A, Maf1-4A) progressively enhanced basal repression of tRNA in actively proliferating cells and attenuated amino acid-induced tRNA transcription. Gene alignment revealed conservation of all four Ser/Thr sites in high eukaryotes, further supporting a critical role of these residues in Maf1 function. Interestingly, mTOR inhibition led to an increase in the occupancy of Maf1 on a set of Pol III-dependent genes, with concomitant reduction in the binding of Pol III and Brf1. Unexpectedly, mTORC1 itself was also enriched at the same set of Pol III templates, but this association was not influenced by mTOR inhibitor treatment. Our results highlight a new and unique mode of regulation of Pol III transcription by mTOR and suggest that normalization of Pol III activity may contribute to the therapeutic efficacy of mTOR inhibitors.
Figures

Similar articles
-
mTOR associates with TFIIIC, is found at tRNA and 5S rRNA genes, and targets their repressor Maf1.Proc Natl Acad Sci U S A. 2010 Jun 29;107(26):11823-8. doi: 10.1073/pnas.1005188107. Epub 2010 Jun 11. Proc Natl Acad Sci U S A. 2010. PMID: 20543138 Free PMC article.
-
Covalent small ubiquitin-like modifier (SUMO) modification of Maf1 protein controls RNA polymerase III-dependent transcription repression.J Biol Chem. 2013 Jun 28;288(26):19288-95. doi: 10.1074/jbc.M113.473744. Epub 2013 May 14. J Biol Chem. 2013. PMID: 23673667 Free PMC article.
-
Recovery of RNA polymerase III transcription from the glycerol-repressed state: revisiting the role of protein kinase CK2 in Maf1 phosphoregulation.J Biol Chem. 2012 Aug 31;287(36):30833-41. doi: 10.1074/jbc.M112.378828. Epub 2012 Jul 18. J Biol Chem. 2012. PMID: 22810236 Free PMC article.
-
MAF1: a new target of mTORC1.Biochem Soc Trans. 2011 Apr;39(2):487-91. doi: 10.1042/BST0390487. Biochem Soc Trans. 2011. PMID: 21428925 Review.
-
Beyond regulation of pol III: Role of MAF1 in growth, metabolism, aging and cancer.Biochim Biophys Acta Gene Regul Mech. 2018 Apr;1861(4):338-343. doi: 10.1016/j.bbagrm.2018.01.019. Epub 2018 Jan 31. Biochim Biophys Acta Gene Regul Mech. 2018. PMID: 29407795 Review.
Cited by
-
Regulation of nuclear mTORC1.Mol Cell Oncol. 2021 May 5;8(3):1896348. doi: 10.1080/23723556.2021.1896348. eCollection 2021. Mol Cell Oncol. 2021. PMID: 34027037 Free PMC article.
-
Deptor transcriptionally regulates endoplasmic reticulum homeostasis in multiple myeloma cells.Oncotarget. 2016 Oct 25;7(43):70546-70558. doi: 10.18632/oncotarget.12060. Oncotarget. 2016. PMID: 27655709 Free PMC article.
-
The RNA polymerase III repressor MAF1 is regulated by ubiquitin-dependent proteasome degradation and modulates cancer drug resistance and apoptosis.J Biol Chem. 2019 Dec 13;294(50):19255-19268. doi: 10.1074/jbc.RA119.008849. Epub 2019 Oct 23. J Biol Chem. 2019. PMID: 31645432 Free PMC article.
-
Rheb regulates nuclear mTORC1 activity independent of farnesylation.Cell Chem Biol. 2022 Jun 16;29(6):1037-1045.e4. doi: 10.1016/j.chembiol.2022.02.006. Epub 2022 Mar 15. Cell Chem Biol. 2022. PMID: 35294906 Free PMC article.
-
Zooming in: PAGE-Northern Blot Helps to Analyze Anti-Sense Transcripts Originating from Human rIGS under Transcriptional Stress.Noncoding RNA. 2021 Aug 24;7(3):50. doi: 10.3390/ncrna7030050. Noncoding RNA. 2021. PMID: 34449671 Free PMC article.
References
-
- Alessi D. R., Pearce L. R., García-Martínez J. M. (2009) Sci. Signal. 2, pe27. - PubMed
-
- Dunlop E. A., Tee A. R. (2009) Cell. Signal. 21, 827–835 - PubMed
-
- Guertin D. A., Sabatini D. M. (2007) Cancer Cell 12, 9–22 - PubMed
-
- Ma X. M., Blenis J. (2009) Nat. Rev. Mol. Cell Biol. 10, 307–318 - PubMed
-
- Engelman J. A. (2009) Nat. Rev. Cancer 9, 550–562 - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

