Prokaryotic ubiquitin-like protein provides a two-part degron to Mycobacterium proteasome substrates
- PMID: 20233925
- PMCID: PMC2876485
- DOI: 10.1128/JB.01639-09
Prokaryotic ubiquitin-like protein provides a two-part degron to Mycobacterium proteasome substrates
Abstract
Prokaryotic ubiquitin-like protein (Pup) is a posttranslational modifier that targets proteins for degradation by the mycobacterial proteasome. We show that the disordered amino terminus of Pup is required for degradation, while the helical carboxyl terminus mediates its attachment to proteins. Thus, Pup has distinct regions that either interact with pupylation enzymes or initiate proteasomal degradation.
Figures

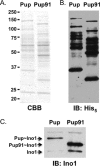
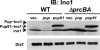
References
-
- Chau, V., J. W. Tobias, A. Bachmair, D. Marriott, D. J. Ecker, D. K. Gonda, and A. Varshavsky. 1989. A multiubiquitin chain is confined to specific lysine in a targeted short-lived protein. Science 243:1576-1583. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

