The SigF regulon in Mycobacterium smegmatis reveals roles in adaptation to stationary phase, heat, and oxidative stress
- PMID: 20233930
- PMCID: PMC2863567
- DOI: 10.1128/JB.00035-10
The SigF regulon in Mycobacterium smegmatis reveals roles in adaptation to stationary phase, heat, and oxidative stress
Abstract
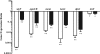
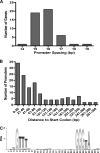
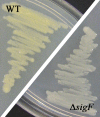
SigF is an alternative sigma factor that is highly conserved among species of the genus Mycobacterium. In this study we identified the SigF regulon in Mycobacterium smegmatis using whole-genome microarray and promoter consensus analyses. In total, 64 genes in exponential phase and 124 genes in stationary phase are SigF dependent (P < 0.01, >2-fold expression change). Our experimental data reveal the SigF-dependent promoter consensus GTTT-N((15-17))-GGGTA for M. smegmatis, and we propose 130 potential genes under direct control of SigF, of which more than 50% exhibited reduced expression in a Delta sigF strain. We previously reported an increased susceptibility of the Delta sigF strain to heat and oxidative stress, and our expression data indicate a molecular basis for these phenotypes. We observed SigF-dependent expression of several genes purportedly involved in oxidative stress defense, namely, a heme-containing catalase, a manganese-containing catalase, a superoxide dismutase, the starvation-induced DNA-protecting protein MsDps1, and the biosynthesis genes for the carotenoid isorenieratene. Our data suggest that SigF regulates the biosynthesis of the thermoprotectant trehalose, as well as an uptake system for osmoregulatory compounds, and this may explain the increased heat susceptibility of the Delta sigF strain. We identified the regulatory proteins SigH3, PhoP, WhiB1, and WhiB4 as possible genes under direct control of SigF and propose four novel anti-sigma factor antagonists that could be involved in the posttranslational regulation of SigF in M. smegmatis. This study emphasizes the importance of this sigma factor for stationary-phase adaptation and stress response in mycobacteria.
Figures
Similar articles
-
Characterization of Mycobacterium smegmatis sigF mutant and its regulon: overexpression of SigF antagonist (MSMEG_1803) in M. smegmatis mimics sigF mutant phenotype, loss of pigmentation, and sensitivity to oxidative stress.Microbiologyopen. 2015 Dec;4(6):896-916. doi: 10.1002/mbo3.288. Epub 2015 Oct 5. Microbiologyopen. 2015. PMID: 26434659 Free PMC article.
-
[Sigma factor F regulates Mycobacterium smegmatis hydrogen peroxide resistance].Wei Sheng Wu Xue Bao. 2012 Nov 4;52(11):1352-9. Wei Sheng Wu Xue Bao. 2012. PMID: 23383506 Chinese.
-
The alternative sigma factor SigF of Mycobacterium smegmatis is required for survival of heat shock, acidic pH and oxidative stress.Microbiology (Reading). 2008 Sep;154(Pt 9):2786-2795. doi: 10.1099/mic.0.2008/018044-0. Microbiology (Reading). 2008. PMID: 18757812
-
Sigma factors and promoters in Corynebacterium glutamicum.J Biotechnol. 2011 Jul 10;154(2-3):101-13. doi: 10.1016/j.jbiotec.2011.01.017. Epub 2011 Jan 26. J Biotechnol. 2011. PMID: 21277915 Review.
-
Elucidating the function of the RpoS regulon.Future Microbiol. 2014;9(4):497-507. doi: 10.2217/fmb.14.9. Future Microbiol. 2014. PMID: 24810349 Review.
Cited by
-
Single START-domain protein Mtsp17 is involved in transcriptional regulation in Mycobacterium smegmatis.PLoS One. 2021 Apr 15;16(4):e0249379. doi: 10.1371/journal.pone.0249379. eCollection 2021. PLoS One. 2021. PMID: 33857164 Free PMC article.
-
A genomic signature and the identification of new sporulation genes.J Bacteriol. 2013 May;195(9):2101-15. doi: 10.1128/JB.02110-12. Epub 2013 Feb 8. J Bacteriol. 2013. PMID: 23396918 Free PMC article.
-
Repression of antibiotic downregulator WblA by AdpA in Streptomyces coelicolor.Appl Environ Microbiol. 2013 Jul;79(13):4159-63. doi: 10.1128/AEM.00546-13. Epub 2013 Apr 19. Appl Environ Microbiol. 2013. PMID: 23603676 Free PMC article.
-
MoaB2, a newly identified transcription factor, binds to σA in Mycobacterium smegmatis.J Bacteriol. 2024 Dec 19;206(12):e0006624. doi: 10.1128/jb.00066-24. Epub 2024 Nov 5. J Bacteriol. 2024. PMID: 39499088 Free PMC article.
-
Tripartite Regulation of the glpFKD Operon Involved in Glycerol Catabolism by GylR, Crp, and SigF in Mycobacterium smegmatis.J Bacteriol. 2019 Nov 20;201(24):e00511-19. doi: 10.1128/JB.00511-19. Print 2019 Dec 15. J Bacteriol. 2019. PMID: 31570530 Free PMC article.
References
-
- Almiron, M., A. J. Link, D. Furlong, and R. Kolter. 1992. A novel DNA-binding protein with regulatory and protective roles in starved Escherichia coli. Genes Dev. 6:2646-2654. - PubMed
-
- Arguelles, J. C. 2000. Physiological roles of trehalose in bacteria and yeasts: a comparative analysis. Arch. Microbiol. 174:217-224. - PubMed
-
- Beaucher, J., S. Rodrigue, P. E. Jacques, I. Smith, R. Brzezinski, and L. Gaudreau. 2002. Novel Mycobacterium tuberculosis anti-sigma factor antagonists control SigmaF activity by distinct mechanisms. Mol. Microbiol. 45:1527-1540. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

