RNA interference-mediated change in protein body morphology and seed opacity through loss of different zein proteins
- PMID: 20237020
- PMCID: PMC2862413
- DOI: 10.1104/pp.110.154690
RNA interference-mediated change in protein body morphology and seed opacity through loss of different zein proteins
Abstract
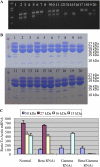
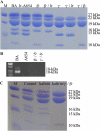
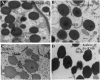
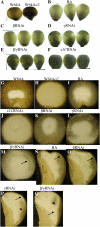
Opaque or nonvitreous phenotypes relate to the seed architecture of maize (Zea mays) and are linked to loci that control the accumulation and proper deposition of storage proteins, called zeins, into specialized organelles in the endosperm, called protein bodies. However, in the absence of null mutants of each type of zein (i.e. alpha, beta, gamma, and delta), the molecular contribution of these proteins to seed architecture remains unclear. Here, a double null mutant for the delta-zeins, the 22-kD alpha-zein, the beta-zein, and the gamma-zein RNA interference (RNAi; designated as z1CRNAi, betaRNAi, and gammaRNAi, respectively) and their combinations have been examined. While the delta-zein double null mutant had negligible effects on protein body formation, the betaRNAi and gammaRNAi alone only cause slight changes. Substantial loss of the 22-kD alpha-zeins by z1CRNAi resulted in protein body budding structures, indicating that a sufficient amount of the 22-kD zeins is necessary for maintenance of a normal protein body shape. Among different mutant combinations, only the combined betaRNAi and gammaRNAi resulted in drastic morphological changes, while other combinations did not. Overexpression of alpha-kafirins, the homologues of the maize 22-kD alpha-zeins in sorghum (Sorghum bicolor), in the beta/gammaRNAi mutant failed to offset the morphological alterations, indicating that beta- and gamma-zeins have redundant and unique functions in the stabilization of protein bodies. Indeed, opacity of the beta/gammaRNAi mutant was caused by incomplete embedding of the starch granules rather than by reducing the vitreous zone.
Figures




Similar articles
-
Gamma-zeins are essential for endosperm modification in quality protein maize.Proc Natl Acad Sci U S A. 2010 Jul 20;107(29):12810-5. doi: 10.1073/pnas.1004721107. Epub 2010 Jul 6. Proc Natl Acad Sci U S A. 2010. PMID: 20615951 Free PMC article.
-
Zein protein interactions, rather than the asymmetric distribution of zein mRNAs on endoplasmic reticulum membranes, influence protein body formation in maize endosperm.Plant Cell. 2002 Mar;14(3):655-72. doi: 10.1105/tpc.010431. Plant Cell. 2002. PMID: 11910012 Free PMC article.
-
Divergent properties of prolamins in wheat and maize.Planta. 2013 Jun;237(6):1465-73. doi: 10.1007/s00425-013-1857-5. Epub 2013 Feb 23. Planta. 2013. PMID: 23435659
-
The regulation of zein biosynthesis in maize endosperm.Theor Appl Genet. 2020 May;133(5):1443-1453. doi: 10.1007/s00122-019-03520-z. Epub 2020 Jan 2. Theor Appl Genet. 2020. PMID: 31897513 Review.
-
Maize opaque mutants are no longer so opaque.Plant Reprod. 2018 Sep;31(3):319-326. doi: 10.1007/s00497-018-0344-3. Epub 2018 Jul 5. Plant Reprod. 2018. PMID: 29978299 Free PMC article. Review.
Cited by
-
Maize Oxalyl-CoA Decarboxylase1 Degrades Oxalate and Affects the Seed Metabolome and Nutritional Quality.Plant Cell. 2018 Oct;30(10):2447-2462. doi: 10.1105/tpc.18.00266. Epub 2018 Sep 10. Plant Cell. 2018. PMID: 30201823 Free PMC article.
-
Towards coeliac-safe bread.Plant Biotechnol J. 2020 Apr;18(4):1056-1065. doi: 10.1111/pbi.13273. Epub 2019 Dec 24. Plant Biotechnol J. 2020. PMID: 31585498 Free PMC article.
-
Comparative transcriptomics reveals the difference in early endosperm development between maize with different amylose contents.PeerJ. 2019 Aug 28;7:e7528. doi: 10.7717/peerj.7528. eCollection 2019. PeerJ. 2019. PMID: 31523504 Free PMC article.
-
Intra-Kernel Reallocation of Proteins in Maize Depends on VP1-Mediated Scutellum Development and Nutrient Assimilation.Plant Cell. 2019 Nov;31(11):2613-2635. doi: 10.1105/tpc.19.00444. Epub 2019 Sep 17. Plant Cell. 2019. PMID: 31530735 Free PMC article.
-
Balancing of sulfur storage in maize seed.BMC Plant Biol. 2012 May 30;12:77. doi: 10.1186/1471-2229-12-77. BMC Plant Biol. 2012. PMID: 22646812 Free PMC article.
References
-
- Chandrashekar A, Mazhar H. (1999) The biochemical basis and implications of grain strength in sorghum and maize. J Cereal Sci 30: 193–207
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

