Cyclin G-associated kinase promotes microtubule outgrowth from chromosomes during spindle assembly
- PMID: 20237935
- PMCID: PMC2919828
- DOI: 10.1007/s00412-010-0267-8
Cyclin G-associated kinase promotes microtubule outgrowth from chromosomes during spindle assembly
Abstract
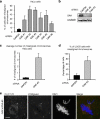
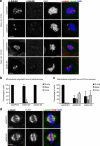
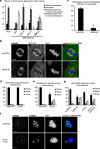
During mitosis, all chromosomes must attach to microtubules of the mitotic spindle to ensure correct chromosome segregation. Microtubule attachment occurs at specialized structures at the centromeric region of chromosomes, called kinetochores. These kinetochores can generate microtubule attachments through capture of centrosome-derived microtubules, but in addition, they can generate microtubules themselves, which are subsequently integrated with centrosome-derived microtubules to form the mitotic spindle. Here, we have performed a large scale RNAi screen and identify cyclin G-associated kinase (GAK) as a novel regulator of microtubule generation at kinetochores/chromatin. This function of GAK requires its C-terminal J-domain, which is essential for clathrin recycling from endocytic vesicles. Consistently, cells lacking GAK show strongly reduced levels of clathrin on the mitotic spindle, and reduction of clathrin levels also inhibits microtubule generation at kinetochores/chromosomes. Finally, we present evidence that association of clathrin with the spindle is promoted by a signal coming from the chromosomes. These results identify a role for GAK and clathrin in microtubule outgrowth from kinetochores/chromosomes and suggest that GAK acts through clathrin to control microtubule outgrowth around chromosomes.
Figures

Similar articles
-
GAK, a regulator of clathrin-mediated membrane traffic, also controls centrosome integrity and chromosome congression.J Cell Sci. 2009 Sep 1;122(Pt 17):3145-52. doi: 10.1242/jcs.052795. Epub 2009 Aug 4. J Cell Sci. 2009. PMID: 19654208
-
The nuclear scaffold protein SAF-A is required for kinetochore-microtubule attachment and contributes to the targeting of Aurora-A to mitotic spindles.J Cell Sci. 2011 Feb 1;124(Pt 3):394-404. doi: 10.1242/jcs.063347. J Cell Sci. 2011. PMID: 21242313
-
The human mitotic checkpoint protein BubR1 regulates chromosome-spindle attachments.Nat Cell Biol. 2005 Jan;7(1):93-8. doi: 10.1038/ncb1208. Epub 2004 Dec 12. Nat Cell Biol. 2005. PMID: 15592459
-
Microtubule capture: a concerted effort.Cell. 2006 Dec 15;127(6):1105-8. doi: 10.1016/j.cell.2006.11.032. Cell. 2006. PMID: 17174890 Review.
-
Chromosome-microtubule interactions during mitosis.Annu Rev Cell Dev Biol. 2002;18:193-219. doi: 10.1146/annurev.cellbio.18.032002.132412. Epub 2002 Apr 2. Annu Rev Cell Dev Biol. 2002. PMID: 12142285 Review.
Cited by
-
The role of clathrin in mitotic spindle organisation.J Cell Sci. 2012 Jan 1;125(Pt 1):19-28. doi: 10.1242/jcs.094607. J Cell Sci. 2012. PMID: 22294613 Free PMC article. Review.
-
Genetic mutations in kinases: a comprehensive review on marketed inhibitors and unexplored targets in Parkinson's disease.Neurol Sci. 2025 Apr;46(4):1509-1524. doi: 10.1007/s10072-024-07970-2. Epub 2025 Jan 6. Neurol Sci. 2025. PMID: 39760821 Review.
-
RNAi screen reveals synthetic lethality between cyclin G-associated kinase and FBXW7 by inducing aberrant mitoses.Br J Cancer. 2017 Sep 26;117(7):954-964. doi: 10.1038/bjc.2017.277. Epub 2017 Aug 22. Br J Cancer. 2017. PMID: 28829765 Free PMC article.
-
Disruption of clathrin-mediated trafficking causes centrosome overduplication and senescence.Traffic. 2014 Jan;15(1):60-77. doi: 10.1111/tra.12132. Epub 2013 Nov 12. Traffic. 2014. PMID: 24138026 Free PMC article.
-
The AP-2 complex interacts with γ-TuRC and regulates the proliferative capacity of neural progenitors.Life Sci Alliance. 2023 Dec 12;7(2):e202302029. doi: 10.26508/lsa.202302029. Print 2024 Feb. Life Sci Alliance. 2023. PMID: 38086550 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

