Single-molecule imaging brings Rad51 nucleoprotein filaments into focus
- PMID: 20299221
- PMCID: PMC2862779
- DOI: 10.1016/j.tcb.2010.02.004
Single-molecule imaging brings Rad51 nucleoprotein filaments into focus
Abstract
The Rad51 protein is essential for DNA repair by homologous recombination. After DNA damage, Rad51 localizes to nuclear foci that represent sites of DNA repair in vivo. In vitro, Rad51 self-assembles on single- or double-stranded DNA to form a nucleoprotein filament. Recently, the merging of innovative single-molecule techniques with ensemble methods has provided unique insights into the dynamic nature of this filament and its cellular function. The assembly and disassembly of Rad51 nucleoprotein filaments is seen to be regulated by recombination accessory proteins. In this regard, the BRC repeats of the BRCA2 protein were shown to modulate the DNA binding selectivity of Rad51. Furthermore, single-molecule studies explained the need for a DNA translocase, Rad54 protein, in the disassembly of Rad51 double-stranded DNA filaments.
Copyright 2010 Elsevier Ltd. All rights reserved.
Figures
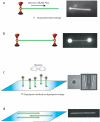
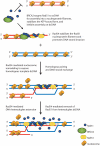
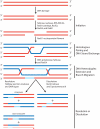
References
-
- Spies M, Kowalczykowski SC. Homologous recombination by RecBCD and RecF pathways. In: Higgins NP, editor. The Bacterial Chromosome. ASM Press; 2005. pp. 389–403.
-
- Lisby M, et al. Choreography of the DNA damage response: spatiotemporal relationships among checkpoint and repair proteins. Cell. 2004;118:699–713. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

