Deletion of the alpha-arrestin protein Txnip in mice promotes adiposity and adipogenesis while preserving insulin sensitivity
- PMID: 20299477
- PMCID: PMC2874703
- DOI: 10.2337/db09-1212
Deletion of the alpha-arrestin protein Txnip in mice promotes adiposity and adipogenesis while preserving insulin sensitivity
Abstract
Objective: Thioredoxin interacting protein (Txnip), a regulator of cellular oxidative stress, is induced by hyperglycemia and inhibits glucose uptake into fat and muscle, suggesting a role for Txnip in type 2 diabetes pathogenesis. Here, we tested the hypothesis that Txnip-null (knockout) mice are protected from insulin resistance induced by a high-fat diet.
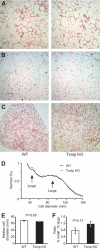
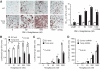
Research design and methods: Txnip gene-deleted (knockout) mice and age-matched wild-type littermate control mice were maintained on a standard chow diet or subjected to 4 weeks of high-fat feeding. Mice were assessed for body composition, fat development, energy balance, and insulin responsiveness. Adipogenesis was measured from ex vivo fat preparations, and in mouse embryonic fibroblasts (MEFs) and 3T3-L1 preadipocytes after forced manipulation of Txnip expression.
Results: Txnip knockout mice gained significantly more adipose mass than controls due to a primary increase in both calorie consumption and adipogenesis. Despite increased fat mass, Txnip knockout mice were markedly more insulin sensitive than controls, and augmented glucose transport was identified in both adipose and skeletal muscle. RNA interference gene-silenced preadipocytes and Txnip(-/-) MEFs were markedly adipogenic, whereas Txnip overexpression impaired adipocyte differentiation. As increased adipogenesis and insulin sensitivity suggested aspects of augmented peroxisome proliferator-activated receptor-gamma (PPARgamma) response, we investigated Txnip's regulation of PPARgamma function; manipulation of Txnip expression directly regulated PPARgamma expression and activity.
Conclusions: Txnip deletion promotes adiposity in the face of high-fat caloric excess; however, loss of this alpha-arrestin protein simultaneously enhances insulin responsiveness in fat and skeletal muscle, revealing Txnip as a novel mediator of insulin resistance and a regulator of adipogenesis.
Figures




References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

