p53 and ARF: unexpected players in autophagy
- PMID: 20303758
- PMCID: PMC2891045
- DOI: 10.1016/j.tcb.2010.02.007
p53 and ARF: unexpected players in autophagy
Abstract
p53 and ARF are well-established tumor-suppressor proteins that function together in the negative regulation of cancer. Recently, both proteins were found to play surprising roles in autophagy. Autophagy ('self-eating') is a crucial response of eukaryotic cells to metabolic and other stress. During this process, portions of the cytosol are sequestered into characteristic double-membrane vesicles that are delivered to the lysosome for degradation, leading to the release of free amino acids and promoting cell survival. The mechanisms whereby p53 and ARF control autophagy are only now becoming elucidated. An emerging question is whether we can develop metabolic poisons that preferentially destroy tumor cells depending on their reliance on autophagy for survival, and on their p53 and ARF status.
(c) 2010 Elsevier Ltd. All rights reserved.
Figures

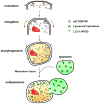


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

