The actinomycin biosynthetic gene cluster of Streptomyces chrysomallus: a genetic hall of mirrors for synthesis of a molecule with mirror symmetry
- PMID: 20304989
- PMCID: PMC2863554
- DOI: 10.1128/JB.01526-09
The actinomycin biosynthetic gene cluster of Streptomyces chrysomallus: a genetic hall of mirrors for synthesis of a molecule with mirror symmetry
Abstract
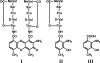
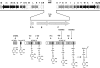
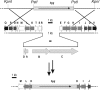
A gene cluster was identified which contains genes involved in the biosynthesis of actinomycin encompassing 50 kb of contiguous DNA on the chromosome of Streptomyces chrysomallus. It contains 28 genes with biosynthetic functions and is bordered on both sides by IS elements. Unprecedentedly, the cluster consists of two large inverted repeats of 11 and 13 genes, respectively, with four nonribosomal peptide synthetase genes in the middle. Nine genes in each repeat have counterparts in the other, in the same arrangement but in the opposite orientation, suggesting an inverse duplication of one of the arms during the evolution of the gene cluster. All of the genes appear to be organized into operons, each corresponding to a functional section of actinomycin biosynthesis, such as peptide assembly, regulation, resistance, and biosynthesis of the precursor of the actinomycin chromophore 4-methyl-3-hydroxyanthranilic acid (4-MHA). For 4-MHA synthesis, functional analysis revealed genes that encode pathway-specific isoforms of tryptophan dioxygenase, kynurenine formamidase, and hydroxykynureninase, which are distinct from the corresponding enzyme activities of cellular tryptophan catabolism in their regulation and in part in their substrate specificity. Phylogenetic analysis indicates that the pathway-specific tryptophan metabolism in Streptomyces most probably evolved divergently from the normal pathway of tryptophan catabolism to provide an extra or independent supply of building blocks for the synthesis of tryptophan-derived secondary metabolites.
Figures
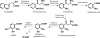
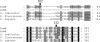



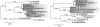

References
-
- Bentley, S. D., K. F. Chater, A. M. Cerdeño-Tárraga, G. L. Challis, N. R. Thomson, K. D. James, D. E. Harris, M. A. Quail, H. Kieser, D. Harper, A. Bateman, S. Brown, G. Chandra, C. W. Chen, M. Collins, A. Cronin, A. Fraser, A. Goble, J. Hidalgo, T. Hornsby, S. Howarth, C. H. Huang, T. Kieser, L. Larke, L. Murphy, K. Oliver, S. O'Neil, E. Rabbinowitsch, M. A. Rajandream, K. Rutherford, S. Rutter, K. Seeger, D. Saunders, S. Sharp, R. Squares, S. Squares, K. Taylor, T. Warren, A. Wietzorrek, J. Woodward, B. G. Barrell, J. Parkhill, and D. A. Hopwood. 2002. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 417:141-147. - PubMed
-
- Bitzer, J., M. Streibel, H. J. Langer, and S. Grond. 2009. First Y-type actinomycins from Streptomyces with divergent structure-activity relationships for antibacterial and cytotoxic properties. Org. Biomol. Chem. 7:444-450. - PubMed
-
- Blondelet-Rouault, M. H., J. Weiser, A. Lebrihi, P. Branny, and J. L. Pernodet. 1997. Antibiotic resistance gene cassettes derived from the omega interposon for use in E. coli and Streptomyces. Gene 190:315-317. - PubMed
-
- Bradford, M. M. 1976. A rapid and sensitive method for the quantitation of microorganism quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72:248. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

