An ancient anion-binding structural module in RNA and DNA helicases
- PMID: 20310069
- PMCID: PMC7610952
- DOI: 10.1002/prot.22704
An ancient anion-binding structural module in RNA and DNA helicases
Abstract
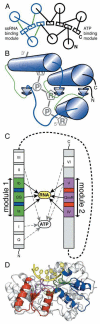
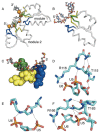
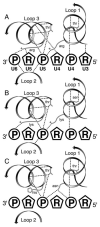
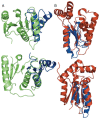
RNA and DNA helicases manipulate or translocate along single strands of nucleic acids by grasping them using a conserved structural motif. We have examined the available crystal structures of helicases of the two principal superfamilies, SF1 and SF2, and observed that the most conserved interactions with the nucleic acid occur between the phosphosugar backbone of a trinucleotide and the three strand-helix loops within a (beta-strand/alpha-helix)(3) structural module. At the first and third loops is a conserved hydrogen-bonded feature called a thr-motif, often seen at alpha-helical N-termini, with the threonine as the N-cap residue. These loops can be aligned with few insertions or deletions, and their main chain atoms are structurally congruent amongst the family members and between the two modules found as tandem pairs in all SF1 and SF2 proteins. The other highly conserved interactions with nucleic acid involve main chain NH groups, often at the helical N-termini, interacting with phosphate groups. We comment on how the sequence motifs that are commonly used to identify helicases map to locations on the module and discuss the implications of the conserved orientation of nucleic acid on the surface of the module for directional stepping along DNA or RNA.
Figures

References
-
- Silverman E, Edwalds-Gilbert G, Lin R-L. DExD/H-box proteins and their partners: helping RNA helicases unwind. Gene. 2003;312:1–16. - PubMed
-
- Singleton MR, Dillingham MS, Wigley DS. Structure and mechanism of helicases and nucleic acid translocases. Ann Rev Biochem. 2007;76:23–50. - PubMed
-
- Pyle A-M. Translocation and unwinding mechanisms of RNA and DNA helicases. Ann Rev Biophys. 2008;37:317–336. - PubMed
-
- Cordin O, Banroques J, Tanner NK, Linder P. Review: the DEADbox protein family of RNA helicases. Gene. 2006;367:17–37. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

