Mapping of drebrin binding site on F-actin
- PMID: 20347847
- PMCID: PMC2866048
- DOI: 10.1016/j.jmb.2010.03.039
Mapping of drebrin binding site on F-actin
Abstract
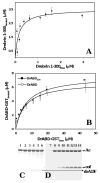
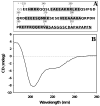
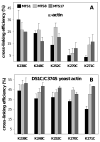
Drebrin is a filament-binding protein involved in organizing the dendritic pool of actin. Previous in vivo studies identified the actin-binding domain of drebrin (DrABD), which causes the same rearrangements in the cytoskeleton as the full-length protein. Site-directed mutagenesis, electron microscopic reconstruction, and chemical cross-linking combined with mass spectrometry analysis were employed here to map the DrABD binding interface on actin filaments. DrABD could be simultaneously attached to two adjacent actin protomers using the combination of 2-iminothiolane (Traut's reagent) and MTS1 [1,1-methanediyl bis(methanethiosulfonate)]. Site-directed mutagenesis combined with chemical cross-linking revealed that residue 238 of DrABD is located within 5.4 A from C374 of actin protomer 1 and that native cysteine 308 of drebrin is near C374 of actin protomer 2. Mass spectrometry analysis revealed that a zero-length cross-linker, 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide, can link the N-terminal G-S extension of the recombinant DrABD to E99 and/or E100 on actin. Efficient cross-linking of drebrin residues 238, 248, 252, 270, and 271 to actin residue 51 was achieved with reagents of different lengths (5.4-19 A). These results suggest that the "core" DrABD is centered on actin subdomain 2 and may adopt a folded conformation upon binding to F-actin. The results of electron microscopic reconstruction, which are in a good agreement with the cross-linking data, revealed polymorphism in DrABD binding to F-actin and suggested the existence of two binding sites. These results provide new structural insight into the previously observed competition between drebrin and several other F-actin-binding proteins.
(c) 2010 Elsevier Ltd. All rights reserved.
Figures
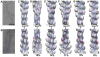
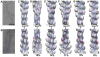




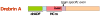
Similar articles
-
Cross-linking constraints on F-actin structure.J Mol Biol. 2000 Jun 2;299(2):421-9. doi: 10.1006/jmbi.2000.3727. J Mol Biol. 2000. PMID: 10860749
-
F-actin structure destabilization and DNase I binding loop: fluctuations mutational cross-linking and electron microscopy analysis of loop states and effects on F-actin.J Mol Biol. 2010 Jan 22;395(3):544-57. doi: 10.1016/j.jmb.2009.11.001. Epub 2009 Nov 6. J Mol Biol. 2010. PMID: 19900461 Free PMC article.
-
Mapping the cofilin binding site on yeast G-actin by chemical cross-linking.J Mol Biol. 2008 Mar 21;377(2):395-409. doi: 10.1016/j.jmb.2007.12.073. Epub 2008 Jan 5. J Mol Biol. 2008. PMID: 18258262 Free PMC article.
-
Biochemistry of Drebrin and Its Binding to Actin Filaments.Adv Exp Med Biol. 2017;1006:37-47. doi: 10.1007/978-4-431-56550-5_3. Adv Exp Med Biol. 2017. PMID: 28865013 Review.
-
Making of a Synapse: Recurrent Roles of Drebrin A at Excitatory Synapses Throughout Life.Adv Exp Med Biol. 2017;1006:119-139. doi: 10.1007/978-4-431-56550-5_8. Adv Exp Med Biol. 2017. PMID: 28865018 Review.
Cited by
-
Actin binding proteins and spermiogenesis: Some unexpected findings.Spermatogenesis. 2011 Apr;1(2):99-104. doi: 10.4161/spmg.1.2.16913. Spermatogenesis. 2011. PMID: 22319657 Free PMC article.
-
Actin-binding protein drebrin E is involved in junction dynamics during spermatogenesis.Spermatogenesis. 2011 Apr-Jun;1(2):123-136. doi: 10.4161/spmg.1.2.16393. Spermatogenesis. 2011. PMID: 22319661 Free PMC article.
-
Biased localization of actin binding proteins by actin filament conformation.Nat Commun. 2020 Nov 25;11(1):5973. doi: 10.1038/s41467-020-19768-9. Nat Commun. 2020. PMID: 33239610 Free PMC article.
-
Cryo-EM reveals different coronin binding modes for ADP- and ADP-BeFx actin filaments.Nat Struct Mol Biol. 2014 Dec;21(12):1075-81. doi: 10.1038/nsmb.2907. Epub 2014 Nov 2. Nat Struct Mol Biol. 2014. PMID: 25362487 Free PMC article.
-
Septin 9 Exhibits Polymorphic Binding to F-Actin and Inhibits Myosin and Cofilin Activity.J Mol Biol. 2015 Oct 9;427(20):3273-3284. doi: 10.1016/j.jmb.2015.07.026. Epub 2015 Aug 19. J Mol Biol. 2015. PMID: 26297986 Free PMC article.
References
-
- Sekino Y, Kojima N, Shirao T. Role of actin cytoskeleton in dendritic spine morphogenesis. Neurochem Int. 2007;51:92–104. - PubMed
-
- Kojima N, Shirao T. Synaptic dysfunction and disruption of postsynaptic drebrin-actin complex: A study of neurological disorders accompanied by cognitive deficits. Neurosci Res. 2007;58:1–5. - PubMed
-
- Peitsch WK, Hofmann I, Pratzel S, Grund C, Kuhn C, Moll I, Langbein L, Franke WW. Drebrin particles: components in the ensemble of proteins regulating actin dynamics of lamellipodia and filopodia. Eur J Cell Biol. 2001;80:567–579. - PubMed
-
- Peitsch WK, Hofmann I, Endlich N, Pratzel S, Kuhn C, Spring H, Grone HJ, Kriz W, Franke WW. Cell biological and biochemical characterization of drebrin complexes in mesangial cells and podocytes of renal glomeruli. J Am Soc Nephrol. 2003;14:1452–1463. - PubMed
-
- Grenklo S, Hillberg L, Zhao Rathje LS, Pinaev G, Schutt C, Lindberg U. Tropomyosin assembly intermediates in the control of microfilament system turnover. Eur J Cell Biol. 2008;87:905–920. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

