Subsurface cycling of nitrogen and anaerobic aromatic hydrocarbon biodegradation revealed by nucleic Acid and metabolic biomarkers
- PMID: 20348302
- PMCID: PMC2869145
- DOI: 10.1128/AEM.00172-10
Subsurface cycling of nitrogen and anaerobic aromatic hydrocarbon biodegradation revealed by nucleic Acid and metabolic biomarkers
Abstract
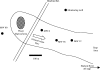
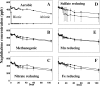
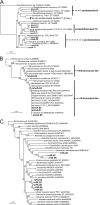
Microbial processes are crucial for ecosystem maintenance, yet documentation of these processes in complex open field sites is challenging. Here we used a multidisciplinary strategy (site geochemistry, laboratory biodegradation assays, and field extraction of molecular biomarkers) to deduce an ongoing linkage between aromatic hydrocarbon biodegradation and nitrogen cycling in a contaminated subsurface site. Three site wells were monitored over a 10-month period, which revealed fluctuating concentrations of nitrate, ammonia, sulfate, sulfide, methane, and other constituents. Biodegradation assays performed under multiple redox conditions indicated that naphthalene metabolism was favored under aerobic conditions. To explore in situ field processes, we measured metabolites of anaerobic naphthalene metabolism and expressed mRNA transcripts selected to document aerobic and anaerobic microbial transformations of ammonia, nitrate, and methylated aromatic contaminants. Gas chromatography-mass spectrometry detection of two carboxylated naphthalene metabolites and transcribed benzylsuccinate synthase, cytochrome c nitrite reductase, and ammonia monooxygenase genes indicated that anaerobic metabolism of aromatic compounds and both dissimilatory nitrate reduction to ammonia (DNRA) and nitrification occurred in situ. These data link formation (via DNRA) and destruction (via nitrification) of ammonia to in situ cycling of nitrogen in this subsurface habitat, where metabolism of aromatic pollutants has led to accumulation of reduced metabolic end products (e.g., ammonia and methane).
Figures
Similar articles
-
Microaerobic biodegradation of aromatic hydrocarbon mixtures: strategies for efficient nitrate and oxygen dosage.Appl Microbiol Biotechnol. 2025 Jan 16;109(1):9. doi: 10.1007/s00253-024-13388-9. Appl Microbiol Biotechnol. 2025. PMID: 39821078 Free PMC article.
-
Bacterial community analysis of shallow groundwater undergoing sequential anaerobic and aerobic chloroethene biotransformation.FEMS Microbiol Ecol. 2007 May;60(2):299-311. doi: 10.1111/j.1574-6941.2007.00290.x. Epub 2007 Mar 26. FEMS Microbiol Ecol. 2007. PMID: 17386036
-
Simultaneous Anaerobic and Aerobic Ammonia and Methane Oxidation under Oxygen Limitation Conditions.Appl Environ Microbiol. 2021 Jun 11;87(13):e0004321. doi: 10.1128/AEM.00043-21. Epub 2021 Jun 11. Appl Environ Microbiol. 2021. PMID: 33893122 Free PMC article.
-
Biodegradation of aromatic compounds under mixed oxygen/denitrifying conditions: a review.J Ind Microbiol Biotechnol. 1997 Feb-Mar;18(2-3):116-30. doi: 10.1038/sj.jim.2900288. J Ind Microbiol Biotechnol. 1997. PMID: 9134760 Review.
-
Anaerobic biodegradation of aromatic hydrocarbons: pathways and prospects.J Mol Microbiol Biotechnol. 2008;15(2-3):93-120. doi: 10.1159/000121324. Epub 2008 Jul 28. J Mol Microbiol Biotechnol. 2008. PMID: 18685265 Review.
Cited by
-
Bacterial nanocellulose production from naphthalene.Microb Biotechnol. 2019 Jul;12(4):662-676. doi: 10.1111/1751-7915.13399. Epub 2019 May 14. Microb Biotechnol. 2019. PMID: 31087504 Free PMC article.
-
Phylogenetic and functional diversity within toluene-degrading, sulphate-reducing consortia enriched from a contaminated aquifer.Microb Ecol. 2014 Aug;68(2):222-34. doi: 10.1007/s00248-014-0403-8. Epub 2014 Mar 13. Microb Ecol. 2014. PMID: 24623528
-
Identification of biomarker genes to predict biodegradation of 1,4-dioxane.Appl Environ Microbiol. 2014 May;80(10):3209-18. doi: 10.1128/AEM.04162-13. Epub 2014 Mar 14. Appl Environ Microbiol. 2014. PMID: 24632253 Free PMC article.
-
Polyphasic analysis of an Azoarcus-Leptothrix-dominated bacterial biofilm developed on stainless steel surface in a gasoline-contaminated hypoxic groundwater.Environ Sci Pollut Res Int. 2016 May;23(9):9019-35. doi: 10.1007/s11356-016-6128-0. Epub 2016 Jan 29. Environ Sci Pollut Res Int. 2016. PMID: 26825521
-
Naphthalene biodegradation under oxygen-limiting conditions: community dynamics and the relevance of biofilm-forming capacity.Microb Biotechnol. 2017 Nov;10(6):1781-1796. doi: 10.1111/1751-7915.12842. Epub 2017 Aug 25. Microb Biotechnol. 2017. PMID: 28840968 Free PMC article.
References
-
- Anneser, B., F. Einsiedl, R. U. Meckenstock, L. Richters, F. Wisotzky, and C. Griebler. 2008. High-resolution monitoring of biogeochemical gradients in a tar oil-contaminated aquifer. Appl. Geochem. 23:1715-1730.
-
- Bailly, J., L. Fraissinet-Tachet, J.-C. Verner, J.-C. Debaud, M. Lemaire, M. Wesolowski-Louvel, and R. Marmeisse. 2007. Soil eukaryotic functional diversity, a metatranscriptomic approach. ISME J. 1:632-642. - PubMed
-
- Bakermans, C., and E. L. Madsen. 2002. Detection in coal tar waste-contaminated groundwater of mRNA transcripts related to naphthalene dioxygenase by fluorescent in situ hybridization with tyramide signal amplification. J. Microbiol. Methods 50:75-84. - PubMed
-
- Bakermans, C., and E. L. Madsen. 2002. Diversity of 16S rDNA and naphthalene dioxygenase genes from coal-tar-waste-contaminated aquifer waters. Microb. Ecol. 44:95-106. - PubMed
-
- Bakermans, C., A. M. Hohnstock-Ashe, S. Padmanabhan, P. Padmanabhan, and E. L. Madsen. 2002. Geochemical and physiological evidence for mixed aerobic and anaerobic field biodegradation of coal tar waste by subsurface microbial communities. Microb. Ecol. 44:107-117. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources

