Interplay between Clostridium thermocellum family 48 and family 9 cellulases in cellulosomal versus noncellulosomal states
- PMID: 20348303
- PMCID: PMC2869131
- DOI: 10.1128/AEM.00009-10
Interplay between Clostridium thermocellum family 48 and family 9 cellulases in cellulosomal versus noncellulosomal states
Abstract
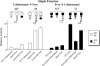
The anaerobic, thermophilic cellulolytic bacterium Clostridium thermocellum is known for its elaborate cellulosome complex, but it also produces a separate free cellulase system. Among the free enzymes, the noncellulosomal enzyme Cel9I is a processive endoglucanase whose sequence and architecture are very similar to those of the cellulosomal enzyme Cel9R; likewise, the noncellulosomal exoglucanase Cel48Y is analogous to the principal cellulosomal enzyme Cel48S. In this study we used the designer cellulosome approach to examine the interplay of prominent cellulosomal and noncellulosomal cellulases from C. thermocellum. Toward this end, we converted the cellulosomal enzymes to noncellulosomal chimeras by swapping the dockerin module of the cellulosomal enzymes with a carbohydrate-binding module from the free enzyme analogues and vice versa. This enabled us to study the importance of the targeting effect of the free enzymes due to their carbohydrate-binding module and the proximity effect for cellulases on the designer cellulosome. C. thermocellum is the only cellulosome-producing bacterium known to express two different glycoside hydrolase family 48 enzymes and thus the only bacterial system that can currently be used for such studies. The different activities with crystalline cellulose were examined, and the results demonstrated that the individual chimeric cellulases were essentially equivalent to the corresponding wild-type analogues. The wild-type cellulases displayed a synergism of about 1.5-fold; the cellulosomal pair acted synergistically when they were converted into free enzymes, whereas the free enzymes acted synergistically mainly in the wild-type state. The targeting effect was found to be the major factor responsible for the elevated activity observed for these specific enzyme combinations, whereas the proximity effect appeared to play a negligible role.
Figures


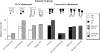
Similar articles
-
Two noncellulosomal cellulases of Clostridium thermocellum, Cel9I and Cel48Y, hydrolyse crystalline cellulose synergistically.FEMS Microbiol Lett. 2007 Mar;268(2):194-201. doi: 10.1111/j.1574-6968.2006.00583.x. Epub 2007 Jan 12. FEMS Microbiol Lett. 2007. PMID: 17227469
-
Stoichiometric Assembly of the Cellulosome Generates Maximum Synergy for the Degradation of Crystalline Cellulose, as Revealed by In Vitro Reconstitution of the Clostridium thermocellum Cellulosome.Appl Environ Microbiol. 2015 Jul;81(14):4756-66. doi: 10.1128/AEM.00772-15. Epub 2015 May 8. Appl Environ Microbiol. 2015. PMID: 25956772 Free PMC article.
-
Incorporation of fungal cellulases in bacterial minicellulosomes yields viable, synergistically acting cellulolytic complexes.Appl Environ Microbiol. 2007 Jun;73(12):3822-32. doi: 10.1128/AEM.00398-07. Epub 2007 Apr 27. Appl Environ Microbiol. 2007. PMID: 17468286 Free PMC article.
-
From cellulosomes to cellulosomics.Chem Rec. 2008;8(6):364-77. doi: 10.1002/tcr.20160. Chem Rec. 2008. PMID: 19107866 Review.
-
The Clostridium cellulovorans cellulosome: an enzyme complex with plant cell wall degrading activity.Chem Rec. 2001;1(1):24-32. doi: 10.1002/1528-0691(2001)1:1<24::AID-TCR5>3.0.CO;2-W. Chem Rec. 2001. PMID: 11893054 Review.
Cited by
-
Metatranscriptomic analysis of lignocellulolytic microbial communities involved in high-solids decomposition of rice straw.Biotechnol Biofuels. 2014 Dec 31;7(1):495. doi: 10.1186/s13068-014-0180-0. eCollection 2014. Biotechnol Biofuels. 2014. PMID: 25648696 Free PMC article.
-
Enhancement of cellulosome-mediated deconstruction of cellulose by improving enzyme thermostability.Biotechnol Biofuels. 2016 Aug 4;9:164. doi: 10.1186/s13068-016-0577-z. eCollection 2016. Biotechnol Biofuels. 2016. PMID: 27493686 Free PMC article.
-
Sequence, structure, and evolution of cellulases in glycoside hydrolase family 48.J Biol Chem. 2012 Nov 30;287(49):41068-77. doi: 10.1074/jbc.M112.405720. Epub 2012 Oct 10. J Biol Chem. 2012. PMID: 23055526 Free PMC article.
-
Improving activity of minicellulosomes by integration of intra- and intermolecular synergies.Biotechnol Biofuels. 2013 Aug 30;6(1):126. doi: 10.1186/1754-6834-6-126. Biotechnol Biofuels. 2013. PMID: 23987588 Free PMC article.
-
Characterization of xylan utilization and discovery of a new endoxylanase in Thermoanaerobacterium saccharolyticum through targeted gene deletions.Appl Environ Microbiol. 2012 Dec;78(23):8441-7. doi: 10.1128/AEM.02130-12. Epub 2012 Sep 28. Appl Environ Microbiol. 2012. PMID: 23023741 Free PMC article.
References
-
- Barak, Y., T. Handelsman, D. Nakar, A. Mechaly, R. Lamed, Y. Shoham, and E. A. Bayer. 2005. Matching fusion-protein systems for affinity analysis of two interacting families of proteins: the cohesin-dockerin interaction. J. Mol. Recognit. 18:491-501. - PubMed
-
- Bayer, E. A., J.-P. Belaich, Y. Shoham, and R. Lamed. 2004. The cellulosomes: multi-enzyme machines for degradation of plant cell wall polysaccharides. Annu. Rev. Microbiol. 58:521-554. - PubMed
-
- Bayer, E. A., B. Henrissat, and R. Lamed. 2008. The cellulosome: a natural bacterial strategy to combat biomass recalcitrance, p. 407-426. In M. E. Himmel (ed.), Biomass recalcitrance. Blackwell, London, United Kingdom.
-
- Bayer, E. A., R. Lamed, and M. E. Himmel. 2007. The potential of cellulases and cellulosomes for cellulosic waste management. Curr. Opin. Biotechnol. 18:237-245. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

