An automated pipeline to screen membrane protein 2D crystallization
- PMID: 20349145
- PMCID: PMC3128831
- DOI: 10.1007/s10969-010-9088-5
An automated pipeline to screen membrane protein 2D crystallization
Abstract
Electron crystallography relies on electron cryomicroscopy of two-dimensional (2D) crystals and is particularly well suited for studying the structure of membrane proteins in their native lipid bilayer environment. To obtain 2D crystals from purified membrane proteins, the detergent in a protein-lipid-detergent ternary mixture must be removed, generally by dialysis, under conditions favoring reconstitution into proteoliposomes and formation of well-ordered lattices. To identify these conditions a wide range of parameters such as pH, lipid composition, lipid-to-protein ratio, ionic strength and ligands must be screened in a procedure involving four steps: crystallization, specimen preparation for electron microscopy, image acquisition, and evaluation. Traditionally, these steps have been carried out manually and, as a result, the scope of 2D crystallization trials has been limited. We have therefore developed an automated pipeline to screen the formation of 2D crystals. We employed a 96-well dialysis block for reconstitution of the target protein over a wide range of conditions designed to promote crystallization. A 96-position magnetic platform and a liquid handling robot were used to prepare negatively stained specimens in parallel. Robotic grid insertion into the electron microscope and computerized image acquisition ensures rapid evaluation of the crystallization screen. To date, 38 2D crystallization screens have been conducted for 15 different membrane proteins, totaling over 3000 individual crystallization experiments. Three of these proteins have yielded diffracting 2D crystals. Our automated pipeline outperforms traditional 2D crystallization methods in terms of throughput and reproducibility.
Figures

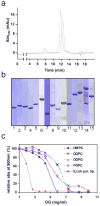



Similar articles
-
Sparse and incomplete factorial matrices to screen membrane protein 2D crystallization.J Struct Biol. 2015 Feb;189(2):123-34. doi: 10.1016/j.jsb.2014.11.008. Epub 2014 Dec 3. J Struct Biol. 2015. PMID: 25478971 Free PMC article.
-
A high-throughput strategy to screen 2D crystallization trials of membrane proteins.J Struct Biol. 2007 Dec;160(3):295-304. doi: 10.1016/j.jsb.2007.09.003. Epub 2007 Sep 14. J Struct Biol. 2007. PMID: 17951070 Free PMC article.
-
The CRACAM Robot: Two-Dimensional Crystallization of Membrane Protein.Methods Mol Biol. 2017;1635:303-316. doi: 10.1007/978-1-4939-7151-0_16. Methods Mol Biol. 2017. PMID: 28755376
-
Electron crystallography of membrane proteins: two-dimensional crystallization and screening by electron microscopy.Methods. 2007 Apr;41(4):417-26. doi: 10.1016/j.ymeth.2006.07.011. Methods. 2007. PMID: 17367714 Review.
-
2D crystallization of membrane proteins: rationales and examples.J Struct Biol. 1998;121(2):162-71. doi: 10.1006/jsbi.1998.3960. J Struct Biol. 1998. PMID: 9615435 Review.
Cited by
-
Automated electron microscopy for evaluating two-dimensional crystallization of membrane proteins.J Struct Biol. 2010 Jul;171(1):102-10. doi: 10.1016/j.jsb.2010.02.018. Epub 2010 Mar 1. J Struct Biol. 2010. PMID: 20197095 Free PMC article.
-
Electron crystallography--the waking beauty of structural biology.Curr Opin Struct Biol. 2012 Aug;22(4):514-9. doi: 10.1016/j.sbi.2012.03.006. Epub 2012 Apr 21. Curr Opin Struct Biol. 2012. PMID: 22525160 Free PMC article. Review.
-
Two-Dimensional Crystallization Procedure, from Protein Expression to Sample Preparation.Biomed Res Int. 2015;2015:693869. doi: 10.1155/2015/693869. Epub 2015 Aug 27. Biomed Res Int. 2015. PMID: 26413539 Free PMC article. Review.
-
Advances in structural and functional analysis of membrane proteins by electron crystallography.Structure. 2011 Oct 12;19(10):1381-93. doi: 10.1016/j.str.2011.09.001. Structure. 2011. PMID: 22000511 Free PMC article. Review.
-
High-throughput methods for electron crystallography.Methods Mol Biol. 2013;955:273-96. doi: 10.1007/978-1-62703-176-9_15. Methods Mol Biol. 2013. PMID: 23132066 Free PMC article.
References
-
- Engelman DM. Lipid bilayer structure in the membrane of Mycoplasma laidlawii. J Mol Biol. 1971;58:153–65. - PubMed
-
- Stevens TJ, Arkin IT. Do more complex organisms have a greater proportion of membrane proteins in their genomes? Proteins. 2000;39:417–20. - PubMed
-
- Planells-Cases R, Jentsch TJ. Chloride channelopathies. Biochim Biophys Acta. 2009;1792:173–89. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

