Discovery of unique lanthionine synthetases reveals new mechanistic and evolutionary insights
- PMID: 20351769
- PMCID: PMC2843593
- DOI: 10.1371/journal.pbio.1000339
Discovery of unique lanthionine synthetases reveals new mechanistic and evolutionary insights
Abstract
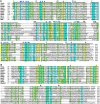
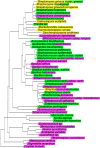
Lantibiotic synthetases are remarkable biocatalysts generating conformationally constrained peptides with a variety of biological activities by repeatedly utilizing two simple posttranslational modification reactions: dehydration of Ser/Thr residues and intramolecular addition of Cys thiols to the resulting dehydro amino acids. Since previously reported lantibiotic synthetases show no apparent homology with any other known protein families, the molecular mechanisms and evolutionary origin of these enzymes are unknown. In this study, we present a novel class of lanthionine synthetases, termed LanL, that consist of three distinct catalytic domains and demonstrate in vitro enzyme activity of a family member from Streptomyces venezuelae. Analysis of individually expressed and purified domains shows that LanL enzymes install dehydroamino acids via phosphorylation of Ser/Thr residues by a protein kinase domain and subsequent elimination of the phosphate by a phosphoSer/Thr lyase domain. The latter has sequence homology with the phosphothreonine lyases found in various pathogenic bacteria that inactivate host mitogen activated protein kinases. A LanC-like cyclase domain then catalyzes the addition of Cys residues to the dehydro amino acids to form the characteristic thioether rings. We propose that LanL enzymes have evolved from stand-alone protein Ser/Thr kinases, phosphoSer/Thr lyases, and enzymes catalyzing thiol alkylation. We also demonstrate that the genes for all three pathways to lanthionine-containing peptides are widespread in Nature. Given the remarkable efficiency of formation of lanthionine-containing polycyclic peptides and the latter's high degree of specificity for their cognate cellular targets, it is perhaps not surprising that (at least) three distinct families of polypeptide sequences have evolved to access this structurally and functionally diverse class of compounds.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




References
-
- Walsh C. T. Washington DC: ASM Press; 2003. Antibiotics: actions, origins, resistance.
-
- Willey J. M, van der Donk W. A. Lantibiotics: peptides of diverse structure and function. Annu Rev Microbiol. 2007;61:477–501. - PubMed
-
- Chatterjee C, Paul M, Xie L, van der Donk W. A. Biosynthesis and mode of action of lantibiotics. Chem Rev. 2005;105:633–684. - PubMed
-
- Breukink E, de Kruijff B. Lipid II as a target for antibiotics. Nat Rev Drug Discov. 2006;5:321–332. - PubMed
-
- Märki F, Hanni E, Fredenhagen A, van Oostrum J. Mode of action of the lanthionine-containing peptide antibiotics duramycin, duramycin B and C, and cinnamycin as indirect inhibitors of phospholipase A2. Biochem Pharmacol. 1991;42:2027–2035. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

