The catecholaminergic polymorphic ventricular tachycardia mutation R33Q disrupts the N-terminal structural motif that regulates reversible calsequestrin polymerization
- PMID: 20353949
- PMCID: PMC2878038
- DOI: 10.1074/jbc.M109.096354
The catecholaminergic polymorphic ventricular tachycardia mutation R33Q disrupts the N-terminal structural motif that regulates reversible calsequestrin polymerization
Abstract
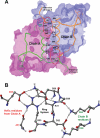
Calsequestrin undergoes dynamic polymerization with increasing calcium concentration by front-to-front dimerization and back-to-back packing, forming wire-shaped structures. A recent finding that point mutation R33Q leads to lethal catecholaminergic polymorphic ventricular tachycardia (CPVT) implies a crucial role for the N terminus. In this study, we demonstrate that this mutation resides in a highly conserved alternately charged residue cluster (DGKDR; cluster 1) in the N-terminal end of calsequestrin. We further show that this cluster configures itself as a ring system and that the dipolar arrangement within the cluster brings about a critical conformational flip of Lys(31)-Asp(32) essential for dimer stabilization by formation of a H-bond network. We additionally show that Ca(2+)-induced calsequestrin aggregation is nonlinear and reversible and can regain the native conformation by Ca(2+) chelation with EGTA. This study suggests that cluster 1 works as a molecular switch and governs the bidirectional transition between the CASQ2 monomer and dimer. We further demonstrate that mutations disrupting the alternating charge pattern of the cluster, including R33Q, impair Ca(2+)-CASQ2 interaction, leading to altered polymerization-depolymerization dynamics. This study provides new mechanistic insight into the functional effects of the R33Q mutation and its potential role in CPVT.
Figures





Similar articles
-
Catecholaminergic polymorphic ventricular tachycardia-related mutations R33Q and L167H alter calcium sensitivity of human cardiac calsequestrin.Biochem J. 2008 Jul 15;413(2):291-303. doi: 10.1042/BJ20080163. Biochem J. 2008. PMID: 18399795
-
Single delivery of an adeno-associated viral construct to transfer the CASQ2 gene to knock-in mice affected by catecholaminergic polymorphic ventricular tachycardia is able to cure the disease from birth to advanced age.Circulation. 2014 Jun 24;129(25):2673-81. doi: 10.1161/CIRCULATIONAHA.113.006901. Epub 2014 Jun 2. Circulation. 2014. PMID: 24888331
-
Probing cationic selectivity of cardiac calsequestrin and its CPVT mutants.Biochem J. 2011 Apr 15;435(2):391-9. doi: 10.1042/BJ20101771. Biochem J. 2011. PMID: 21265816
-
Calsequestrin mutations and catecholaminergic polymorphic ventricular tachycardia.Pediatr Cardiol. 2012 Aug;33(6):959-67. doi: 10.1007/s00246-012-0256-1. Epub 2012 Mar 16. Pediatr Cardiol. 2012. PMID: 22421959 Free PMC article. Review.
-
Calsequestrin 2 and arrhythmias.Am J Physiol Heart Circ Physiol. 2012 Mar 15;302(6):H1250-60. doi: 10.1152/ajpheart.00779.2011. Epub 2011 Dec 23. Am J Physiol Heart Circ Physiol. 2012. PMID: 22198169 Free PMC article. Review.
Cited by
-
Phylogenetic and biochemical analysis of calsequestrin structure and association of its variants with cardiac disorders.Sci Rep. 2020 Oct 22;10(1):18115. doi: 10.1038/s41598-020-75097-3. Sci Rep. 2020. PMID: 33093545 Free PMC article.
-
Inherited dysfunction of sarcoplasmic reticulum Ca2+ handling and arrhythmogenesis.Circ Res. 2011 Apr 1;108(7):871-83. doi: 10.1161/CIRCRESAHA.110.226845. Circ Res. 2011. PMID: 21454795 Free PMC article. Review.
-
The structure of a calsequestrin filament reveals mechanisms of familial arrhythmia.Nat Struct Mol Biol. 2020 Dec;27(12):1142-1151. doi: 10.1038/s41594-020-0510-9. Epub 2020 Oct 12. Nat Struct Mol Biol. 2020. PMID: 33046906 Free PMC article.
-
An International Multicenter Evaluation of Inheritance Patterns, Arrhythmic Risks, and Underlying Mechanisms of CASQ2-Catecholaminergic Polymorphic Ventricular Tachycardia.Circulation. 2020 Sep 8;142(10):932-947. doi: 10.1161/CIRCULATIONAHA.120.045723. Epub 2020 Jul 22. Circulation. 2020. PMID: 32693635 Free PMC article. Clinical Trial.
-
Identification of calcium binding sites on calsequestrin 1 and their implications for polymerization.Mol Biosyst. 2013 Jul;9(7):1949-57. doi: 10.1039/c3mb25588c. Epub 2013 Apr 29. Mol Biosyst. 2013. PMID: 23629537 Free PMC article.
References
-
- Bers D. M. (1997) Basic Res. Cardiol. 92, Suppl. 1, 1–10 - PubMed
-
- MacLennan D. H., Campbell K. P., Reithmeier R. A. F. (1983) in Calcium and Cell Function (Cheung W. Y. ed) Vol. IV, pp. 151–173, Academic Press, Inc., New York
-
- Slupsky J. R., Ohnishi M., Carpenter M. R., Reithmeier R. A. (1987) Biochemistry 26, 6539–6544 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

