In vivo trans-specific gene silencing in fungal cells by in planta expression of a double-stranded RNA
- PMID: 20356372
- PMCID: PMC2907587
- DOI: 10.1186/1741-7007-8-27
In vivo trans-specific gene silencing in fungal cells by in planta expression of a double-stranded RNA
Abstract
Background: Self-complementary RNA transcripts form a double-stranded RNA (dsRNA) that triggers a sequence-specific mRNA degradation, in a process known as RNA interference (RNAi), leading to gene silencing. In vascular plants, RNAi molecules trafficking occur between cells and systemically throughout the plant. RNAi signals can spread systemically throughout a plant, even across graft junctions from transgenic to non-transgenic stocks. There is also a great interest in applying RNAi to pathogenic fungi. Specific inhibition of gene expression by RNAi has been shown to be suitable for a multitude of phytopathogenic filamentous fungi. However, double-stranded (ds)RNA/small interfering (si)RNA silencing effect has not been observed in vivo.
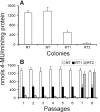
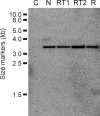
Results: This study demonstrates for the first time the in vivo interference phenomenon in the pathogenic fungus Fusarium verticillioides, in which expression of an individual fungal transgene was specifically abolished by inoculating mycelial cells in transgenic tobacco plants engineered to express siRNAs from a dsRNA corresponding to the particular transgene.
Conclusion: The results provide a powerful tool for further studies on molecular plant-microbe and symbiotic interactions. From a biotechnological perspective, silencing of fungal genes by generating siRNAs in the host provides a novel strategy for the development of broad fungi-resistance strategies in plants and other organisms.
Figures



Similar articles
-
Exogenous application of double-stranded RNA molecules from TMV p126 and CP genes confers resistance against TMV in tobacco.Planta. 2016 Oct;244(4):961-9. doi: 10.1007/s00425-016-2567-6. Epub 2016 Jul 25. Planta. 2016. PMID: 27456838
-
Host-delivered RNAi: an effective strategy to silence genes in plant parasitic nematodes.Planta. 2007 Nov;226(6):1525-33. doi: 10.1007/s00425-007-0588-x. Epub 2007 Jul 25. Planta. 2007. PMID: 17653759
-
Silencing efficiency of dsRNA fragments targeting Fusarium graminearum TRI6 and patterns of small interfering RNA associated with reduced virulence and mycotoxin production.PLoS One. 2018 Aug 30;13(8):e0202798. doi: 10.1371/journal.pone.0202798. eCollection 2018. PLoS One. 2018. PMID: 30161200 Free PMC article.
-
Post-transcriptional gene silencing: Basic concepts and applications.J Biosci. 2020;45:128. J Biosci. 2020. PMID: 33184244 Review.
-
Exogenous RNAs for Gene Regulation and Plant Resistance.Int J Mol Sci. 2019 May 8;20(9):2282. doi: 10.3390/ijms20092282. Int J Mol Sci. 2019. PMID: 31072065 Free PMC article. Review.
Cited by
-
Host Induced Gene Silencing Targeting Aspergillus flavus aflM Reduced Aflatoxin Contamination in Transgenic Maize Under Field Conditions.Front Microbiol. 2020 Apr 28;11:754. doi: 10.3389/fmicb.2020.00754. eCollection 2020. Front Microbiol. 2020. PMID: 32411110 Free PMC article.
-
Host-induced silencing of essential genes in Puccinia triticina through transgenic expression of RNAi sequences reduces severity of leaf rust infection in wheat.Plant Biotechnol J. 2018 May;16(5):1013-1023. doi: 10.1111/pbi.12845. Epub 2017 Dec 15. Plant Biotechnol J. 2018. PMID: 28941315 Free PMC article.
-
Functional characterization of calcineurin homologs PsCNA1/PsCNB1 in Puccinia striiformis f. sp. tritici using a host-induced RNAi system.PLoS One. 2012;7(11):e49262. doi: 10.1371/journal.pone.0049262. Epub 2012 Nov 6. PLoS One. 2012. PMID: 23139840 Free PMC article.
-
Role of Nrf2 signaling pathway in the radiation tolerance of patients with head and neck squamous cell carcinoma: an in vivo and in vitro study.Onco Targets Ther. 2017 Mar 23;10:1809-1819. doi: 10.2147/OTT.S122803. eCollection 2017. Onco Targets Ther. 2017. Retraction in: Onco Targets Ther. 2022 Jul 28;15:827-828. doi: 10.2147/OTT.S383774. PMID: 28367064 Free PMC article. Retracted.
-
Reducing production of fumonisin mycotoxins in Fusarium verticillioides by RNA interference.Mycotoxin Res. 2018 Mar;34(1):29-37. doi: 10.1007/s12550-017-0296-8. Epub 2017 Nov 22. Mycotoxin Res. 2018. PMID: 29164518
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

