A domesticated piggyBac transposase plays key roles in heterochromatin dynamics and DNA cleavage during programmed DNA deletion in Tetrahymena thermophila
- PMID: 20357003
- PMCID: PMC2869380
- DOI: 10.1091/mbc.e09-12-1079
A domesticated piggyBac transposase plays key roles in heterochromatin dynamics and DNA cleavage during programmed DNA deletion in Tetrahymena thermophila
Abstract
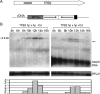
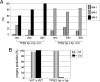
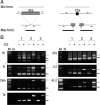
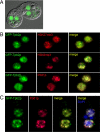
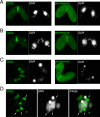
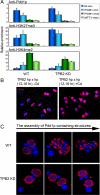
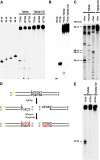
Transposons comprise large fractions of eukaryotic genomes and provide genetic reservoirs for the evolution of new cellular functions. We identified TPB2, a homolog of the piggyBac transposase gene that is required for programmed DNA deletion in Tetrahymena. TPB2 was expressed exclusively during the time of DNA excision, and its encoded protein Tpb2p was localized in DNA elimination heterochromatin structures. Notably, silencing of TPB2 by RNAi disrupts the final assembly of these heterochromatin structures and prevents DNA deletion to occur. In vitro studies revealed that Tpb2p is an endonuclease that produces double-strand breaks with four-base 5' protruding ends, similar to the ends generated during DNA deletion. These findings suggest that Tpb2p plays a key role in the assembly of specialized DNA elimination chromatin architectures and is likely responsible for the DNA cleavage step of programmed DNA deletion.
Figures

Similar articles
-
A domesticated PiggyBac transposase interacts with heterochromatin and catalyzes reproducible DNA elimination in Tetrahymena.PLoS Genet. 2013;9(12):e1004032. doi: 10.1371/journal.pgen.1004032. Epub 2013 Dec 12. PLoS Genet. 2013. PMID: 24348275 Free PMC article.
-
The taming of the shrew: Regulation of a catalytically active domesticated transposase.Mob Genet Elements. 2014 May 27;4:e29383. doi: 10.4161/mge.29383. eCollection 2014. Mob Genet Elements. 2014. PMID: 25054083 Free PMC article.
-
A germline-limited piggyBac transposase gene is required for precise excision in Tetrahymena genome rearrangement.Nucleic Acids Res. 2017 Sep 19;45(16):9481-9502. doi: 10.1093/nar/gkx652. Nucleic Acids Res. 2017. PMID: 28934495 Free PMC article.
-
Programmed Genome Rearrangements in Tetrahymena.Microbiol Spectr. 2014 Dec;2(6). doi: 10.1128/microbiolspec.MDNA3-0012-2014. Microbiol Spectr. 2014. PMID: 26104448 Review.
-
Dynamic nuclear reorganization during genome remodeling of Tetrahymena.Biochim Biophys Acta. 2008 Nov;1783(11):2130-6. doi: 10.1016/j.bbamcr.2008.07.012. Epub 2008 Jul 28. Biochim Biophys Acta. 2008. PMID: 18706458 Free PMC article. Review.
Cited by
-
Chromosome fusions triggered by noncoding RNA.RNA Biol. 2017 May 4;14(5):620-631. doi: 10.1080/15476286.2016.1195940. Epub 2016 Jun 7. RNA Biol. 2017. PMID: 27267579 Free PMC article.
-
Identification of RAG-like transposons in protostomes suggests their ancient bilaterian origin.Mob DNA. 2020 May 6;11:17. doi: 10.1186/s13100-020-00214-y. eCollection 2020. Mob DNA. 2020. PMID: 32399063 Free PMC article.
-
DNA rearrangements directed by non-coding RNAs in ciliates.Wiley Interdiscip Rev RNA. 2010 Nov-Dec;1(3):376-87. doi: 10.1002/wrna.34. Wiley Interdiscip Rev RNA. 2010. PMID: 21956937 Free PMC article. Review.
-
Developmentally programmed, RNA-directed genome rearrangement in Tetrahymena.Dev Growth Differ. 2012 Jan;54(1):108-19. doi: 10.1111/j.1440-169X.2011.01305.x. Epub 2011 Nov 22. Dev Growth Differ. 2012. PMID: 22103557 Free PMC article. Review.
-
Chromatin-associated ncRNA activities.Chromosome Res. 2013 Dec;21(6-7):627-41. doi: 10.1007/s10577-013-9390-8. Chromosome Res. 2013. PMID: 24249576 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

