Autologous neutralizing antibodies to the transmitted/founder viruses emerge late after simian immunodeficiency virus SIVmac251 infection of rhesus monkeys
- PMID: 20357097
- PMCID: PMC2876635
- DOI: 10.1128/JVI.02741-09
Autologous neutralizing antibodies to the transmitted/founder viruses emerge late after simian immunodeficiency virus SIVmac251 infection of rhesus monkeys
Abstract
While the simian immunodeficiency virus (SIV)-infected rhesus monkey is an important animal model for human immunodeficiency virus type 1 (HIV-1) infection of humans, much remains to be learned about the evolution of the humoral immune response in this model. In HIV-1 infection, autologous neutralizing antibodies emerge 2 to 3 months after infection. However, the ontogeny of the SIV-specific neutralizing antibody response in mucosally infected animals has not been defined. We characterized the kinetics of the autologous neutralizing antibody response to the transmitted/founder SIVmac251 using a pseudovirion-based TZM-bl cell assay and monitored env sequence evolution using single-genome amplification in four rhesus animals that were infected via intrarectal inoculations. We show that the SIVmac251 founder viruses induced neutralizing antibodies at 5 to 8 months after infection. Despite their slow emergence and low titers, these neutralizing antibodies selected for escape mutants that harbored substitutions and deletions in variable region 1 (V1), V2, and V4 of Env. The neutralizing antibody response was initially focused on V4 at 5 to 8 months after infection and then targeted V1/V2 and V4 by 16 months. These findings reveal a striking delay in the development of neutralizing antibodies in SIVmac-infected animals, thus raising questions concerning the suitability of SIVmac251 as a challenge strain to screen AIDS vaccines that elicit neutralizing antibodies as a means to prevent virus acquisition. They also illustrate the capacity of the SIVmac quasispecies to modify antigenic determinants in response to very modest titers of neutralizing antibodies.
Figures


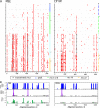



Similar articles
-
Envelope variable region 4 is the first target of neutralizing antibodies in early simian immunodeficiency virus mac251 infection of rhesus monkeys.J Virol. 2012 Jul;86(13):7052-9. doi: 10.1128/JVI.00107-12. Epub 2012 Apr 24. J Virol. 2012. PMID: 22532675 Free PMC article.
-
Limited contribution of mucosal IgA to Simian immunodeficiency virus (SIV)-specific neutralizing antibody response and virus envelope evolution in breast milk of SIV-infected, lactating rhesus monkeys.J Virol. 2010 Aug;84(16):8209-18. doi: 10.1128/JVI.00656-10. Epub 2010 Jun 2. J Virol. 2010. PMID: 20519381 Free PMC article.
-
Breakthrough of SIV strain smE660 challenge in SIV strain mac239-vaccinated rhesus macaques despite potent autologous neutralizing antibody responses.Proc Natl Acad Sci U S A. 2015 Aug 25;112(34):10780-5. doi: 10.1073/pnas.1509731112. Epub 2015 Aug 10. Proc Natl Acad Sci U S A. 2015. PMID: 26261312 Free PMC article.
-
CD4-HIV-1 Envelope Interactions: Critical Insights for the Simian/HIV/Macaque Model.AIDS Res Hum Retroviruses. 2018 Sep;34(9):778-779. doi: 10.1089/AID.2018.0110. Epub 2018 Jul 9. AIDS Res Hum Retroviruses. 2018. PMID: 29886767 Free PMC article. Review.
-
Specificity of the autologous neutralizing antibody response.Curr Opin HIV AIDS. 2009 Sep;4(5):358-63. doi: 10.1097/COH.0b013e32832ea7e8. Curr Opin HIV AIDS. 2009. PMID: 20048698 Free PMC article. Review.
Cited by
-
SIV infection duration largely determines broadening of neutralizing antibody response in macaques.J Clin Invest. 2020 Oct 1;130(10):5413-5424. doi: 10.1172/JCI139123. J Clin Invest. 2020. PMID: 32663192 Free PMC article.
-
The Mtb-HIV syndemic interaction: why treating M. tuberculosis infection may be crucial for HIV-1 eradication.Future Virol. 2020 Feb;15(2):101-125. doi: 10.2217/fvl-2019-0069. Future Virol. 2020. PMID: 32273900 Free PMC article. Review.
-
Depletion of CD4⁺ T cells abrogates post-peak decline of viremia in SIV-infected rhesus macaques.J Clin Invest. 2011 Nov;121(11):4433-45. doi: 10.1172/JCI46023. Epub 2011 Oct 17. J Clin Invest. 2011. PMID: 22005304 Free PMC article.
-
Molecular evolution analysis of the human immunodeficiency virus type 1 envelope in simian/human immunodeficiency virus-infected macaques: implications for challenge dose selection.J Virol. 2011 Oct;85(19):10332-45. doi: 10.1128/JVI.05290-11. Epub 2011 Jul 27. J Virol. 2011. PMID: 21795341 Free PMC article.
-
Early low-titer neutralizing antibodies impede HIV-1 replication and select for virus escape.PLoS Pathog. 2012;8(5):e1002721. doi: 10.1371/journal.ppat.1002721. Epub 2012 May 31. PLoS Pathog. 2012. PMID: 22693447 Free PMC article.
References
-
- Babas, T., B. Belhadj-Jrad, R. Le Grand, D. Dormont, L. Montagnier, and E. Bahraoui. 1995. Specificity and neutralizing capacity of three monoclonal antibodies produced against the envelope glycoprotein of simian immunodeficiency virus isolate 251. Virology 211:339-344. - PubMed
-
- Basavapathruni, A., W. W. Yeh, R. T. Coffey, J. B. Whitney, P. T. Hraber, A. Giri, B. T. Korber, S. S. Rao, G. J. Nabel, J. R. Mascola, M. S. Seaman, and N. L. Letvin. 2010. Envelope vaccination shapes viral envelope evolution following simian immunodeficiency virus infection in rhesus monkeys. J. Virol. 84:953-963. - PMC - PubMed
-
- Bhattacharya, T., M. Daniels, D. Heckerman, B. Foley, N. Frahm, C. Kadie, J. Carlson, K. Yusim, B. McMahon, B. Gaschen, S. Mallal, J. I. Mullins, D. C. Nickle, J. Herbeck, C. Rousseau, G. H. Learn, T. Miura, C. Brander, B. Walker, and B. Korber. 2007. Founder effects in the assessment of HIV polymorphisms and HLA allele associations. Science 315:1583-1586. - PubMed
-
- Blay, W. M., S. Gnanakaran, B. Foley, N. A. Doria-Rose, B. T. Korber, and N. L. Haigwood. 2006. Consistent patterns of change during the divergence of human immunodeficiency virus type 1 envelope from that of the inoculated virus in simian/human immunodeficiency virus-infected macaques. J. Virol. 80:999-1014. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

