The YlmG protein has a conserved function related to the distribution of nucleoids in chloroplasts and cyanobacteria
- PMID: 20359373
- PMCID: PMC2923531
- DOI: 10.1186/1471-2229-10-57
The YlmG protein has a conserved function related to the distribution of nucleoids in chloroplasts and cyanobacteria
Abstract
Background: Reminiscent of their free-living cyanobacterial ancestor, chloroplasts proliferate by division coupled with the partition of nucleoids (DNA-protein complexes). Division of the chloroplast envelope membrane is performed by constriction of the ring structures at the division site. During division, nucleoids also change their shape and are distributed essentially equally to the daughter chloroplasts. Although several components of the envelope division machinery have been identified and characterized, little is known about the molecular components/mechanisms underlying the change of the nucleoid structure.
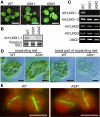
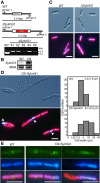
Results: In order to identify new factors that are involved in the chloroplast division, we isolated Arabidopsis thaliana chloroplast division mutants from a pool of random cDNA-overexpressed lines. We found that the overexpression of a previously uncharacterized gene (AtYLMG1-1) of cyanobacterial origin results in the formation of an irregular network of chloroplast nucleoids, along with a defect in chloroplast division. In contrast, knockdown of AtYLMG1-1 resulted in a concentration of the nucleoids into a few large structures, but did not affect chloroplast division. Immunofluorescence microscopy showed that AtYLMG1-1 localizes in small puncta on thylakoid membranes, to which a subset of nucleoids colocalize. In addition, in the cyanobacterium Synechococcus elongates, overexpression and deletion of ylmG also displayed defects in nucleoid structure and cell division.
Conclusions: These results suggest that the proper distribution of nucleoids requires the YlmG protein, and the mechanism is conserved between cyanobacteria and chloroplasts. Given that ylmG exists in a cell division gene cluster downstream of ftsZ in gram-positive bacteria and that ylmG overexpression impaired the chloroplast division, the nucleoid partitioning by YlmG might be related to chloroplast and cyanobacterial division processes.
Figures



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

