Constrained intron structures in a microsporidian
- PMID: 20360213
- PMCID: PMC3809491
- DOI: 10.1093/molbev/msq087
Constrained intron structures in a microsporidian
Abstract
The 2.9-Mbp genome of the microsporidian Encephalitozoon cuniculi is severely reduced and compacted, possessing only 16 known tiny spliceosomal introns. Based on motif and expression data, intron profiles were constructed to screen the genome. Twenty additional introns were predicted and verified, doubling the previous estimate. We further predict that accurate 3' splice site (3'SS) selection is accomplished via a scanning mechanism with specificity achieved by maintaining a constrained variable length between the branch point motif and 3'SS. Only introns in ribosomal protein genes exhibit positional bias, and we hypothesize that splicing could be regulating expression of these genes. The large set of new introns in non-ribosomal protein genes suggests that current models of intron loss are unlikely sufficient to explain the distribution of introns. Together, these results extend our understanding of the role of intron loss in genome evolution and contribute to a novel model for splice site selection.
Figures
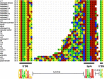


Similar articles
-
Microsporidian Introns Retained against a Background of Genome Reduction: Characterization of an Unusual Set of Introns.Genome Biol Evol. 2019 Jan 1;11(1):263-269. doi: 10.1093/gbe/evy260. Genome Biol Evol. 2019. PMID: 30496512 Free PMC article.
-
A small spliceosomal-type intron occurs in a ribosomal protein gene of the microsporidian Encephalitozoon cuniculi.Mol Biochem Parasitol. 1998 Aug 1;94(2):283-6. doi: 10.1016/s0166-6851(98)00064-4. Mol Biochem Parasitol. 1998. PMID: 9747977 No abstract available.
-
Transcriptome analysis of the parasite Encephalitozoon cuniculi: an in-depth examination of pre-mRNA splicing in a reduced eukaryote.BMC Genomics. 2013 Mar 28;14:207. doi: 10.1186/1471-2164-14-207. BMC Genomics. 2013. PMID: 23537046 Free PMC article.
-
The intronome of budding yeasts.C R Biol. 2011 Aug-Sep;334(8-9):662-70. doi: 10.1016/j.crvi.2011.05.015. Epub 2011 Jul 6. C R Biol. 2011. PMID: 21819948 Review.
-
The microsporidian Encephalitozoon.Bioessays. 2001 Feb;23(2):194-202. doi: 10.1002/1521-1878(200102)23:2<194::AID-BIES1027>3.0.CO;2-D. Bioessays. 2001. PMID: 11169593 Review.
Cited by
-
Analysis of Fungal Genomes Reveals Commonalities of Intron Gain or Loss and Functions in Intron-Poor Species.Mol Biol Evol. 2021 Sep 27;38(10):4166-4186. doi: 10.1093/molbev/msab094. Mol Biol Evol. 2021. PMID: 33772558 Free PMC article.
-
The genome of the obligate intracellular parasite Trachipleistophora hominis: new insights into microsporidian genome dynamics and reductive evolution.PLoS Pathog. 2012;8(10):e1002979. doi: 10.1371/journal.ppat.1002979. Epub 2012 Oct 25. PLoS Pathog. 2012. PMID: 23133373 Free PMC article.
-
The genome of Spraguea lophii and the basis of host-microsporidian interactions.PLoS Genet. 2013;9(8):e1003676. doi: 10.1371/journal.pgen.1003676. Epub 2013 Aug 22. PLoS Genet. 2013. PMID: 23990793 Free PMC article.
-
Contrasting outcomes of genome reduction in mikrocytids and microsporidians.BMC Biol. 2023 Jun 6;21(1):137. doi: 10.1186/s12915-023-01635-w. BMC Biol. 2023. PMID: 37280585 Free PMC article.
-
Evolution and Diversity of Pre-mRNA Splicing in Highly Reduced Nucleomorph Genomes.Genome Biol Evol. 2018 Jun 1;10(6):1573-1583. doi: 10.1093/gbe/evy111. Genome Biol Evol. 2018. PMID: 29860351 Free PMC article.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Miscellaneous

