Hepatitis C virus genomic RNA dimerization is mediated via a kissing complex intermediate
- PMID: 20360391
- PMCID: PMC2856886
- DOI: 10.1261/rna.1960410
Hepatitis C virus genomic RNA dimerization is mediated via a kissing complex intermediate
Abstract
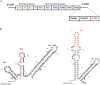
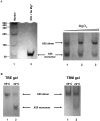
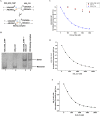
With over 200 million people infected with hepatitis C virus (HCV) worldwide, there is a need for more effective and better-tolerated therapeutic strategies. The HCV genome is a positive-sense; single-stranded RNA encoding a large polyprotein cleaved at multiple sites to produce at least ten proteins, among them an error-prone RNA polymerase that confers a high mutation rate. Despite considerable overall sequence diversity, in the 3'-untranslated region of the HCV genomic RNA there is a 98-nucleotide (nt) sequence named X RNA, the first 55 nt of which (X55 RNA) are 100% conserved among all HCV strains. The X55 region has been suggested to be responsible for in vitro dimerization of the genomic RNA in the presence of the viral core protein, although the mechanism by which this occurs is unknown. In this study, we analyzed the X55 region and characterized the mechanism by which it mediates HCV genomic RNA dimerization. Similar to a mechanism proposed previously for the human immunodeficiency 1 virus (HIV-1) genome, we show that dimerization of the HCV genome involves formation of a kissing complex intermediate, which is converted to a more stable extended duplex conformation in the presence of the core protein. Mutations in the dimer linkage sequence loop sequence that prevent RNA dimerization in vitro significantly reduced but did not completely ablate the ability of HCV RNA to replicate or produce infectious virus in transfected cells.
Figures
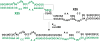


Similar articles
-
Form confers function: Case of the 3'X region of the hepatitis C virus genome.World J Gastroenterol. 2018 Aug 14;24(30):3374-3383. doi: 10.3748/wjg.v24.i30.3374. World J Gastroenterol. 2018. PMID: 30122877 Free PMC article. Review.
-
The hepatitis C virus Core protein is a potent nucleic acid chaperone that directs dimerization of the viral (+) strand RNA in vitro.Nucleic Acids Res. 2004 May 11;32(8):2623-31. doi: 10.1093/nar/gkh579. Print 2004. Nucleic Acids Res. 2004. PMID: 15141033 Free PMC article.
-
Analysis of hepatitis C virus RNA dimerization and core-RNA interactions.Nucleic Acids Res. 2006 May 17;34(9):2618-33. doi: 10.1093/nar/gkl240. Print 2006. Nucleic Acids Res. 2006. PMID: 16707664 Free PMC article.
-
Analysis of the RNA chaperoning activity of the hepatitis C virus core protein on the conserved 3'X region of the viral genome.Nucleic Acids Res. 2012 Mar;40(6):2540-53. doi: 10.1093/nar/gkr1140. Epub 2011 Nov 29. Nucleic Acids Res. 2012. PMID: 22127859 Free PMC article.
-
[Structure and function of the non-coding regions of hepatitis C viral RNA].Postepy Biochem. 2006;52(1):62-71. Postepy Biochem. 2006. PMID: 16869303 Review. Polish.
Cited by
-
An unexpected RNA distal interaction mode found in an essential region of the hepatitis C virus genome.Nucleic Acids Res. 2018 May 4;46(8):4200-4212. doi: 10.1093/nar/gky074. Nucleic Acids Res. 2018. PMID: 29409065 Free PMC article.
-
The conserved 3'X terminal domain of hepatitis C virus genomic RNA forms a two-stem structure that promotes viral RNA dimerization.Nucleic Acids Res. 2015 Sep 30;43(17):8529-39. doi: 10.1093/nar/gkv786. Epub 2015 Aug 3. Nucleic Acids Res. 2015. PMID: 26240378 Free PMC article.
-
Hepatitis C virus RNA: molecular switches mediated by long-range RNA-RNA interactions?Nucleic Acids Res. 2013 Feb 1;41(4):2526-40. doi: 10.1093/nar/gks1318. Epub 2012 Dec 28. Nucleic Acids Res. 2013. PMID: 23275555 Free PMC article.
-
Form confers function: Case of the 3'X region of the hepatitis C virus genome.World J Gastroenterol. 2018 Aug 14;24(30):3374-3383. doi: 10.3748/wjg.v24.i30.3374. World J Gastroenterol. 2018. PMID: 30122877 Free PMC article. Review.
-
Three-dimensional structure of the 3'X-tail of hepatitis C virus RNA in monomeric and dimeric states.RNA. 2017 Sep;23(9):1465-1476. doi: 10.1261/rna.060632.117. Epub 2017 Jun 19. RNA. 2017. PMID: 28630140 Free PMC article.
References
-
- Aagaard L, Rasmussen SV, Mikkelsen JG, Pedersen FS 2004. Efficient replication of full-length murine leukemia viruses modified at the dimer initiation site regions. Virology 318: 360–370 - PubMed
-
- Argaman L, Altuvia S 2000. fhlA repression by OxyS RNA: Kissing complex formation at two sites results in a stable antisense-target RNA complex. J Mol Biol 300: 1101–1112 - PubMed
-
- Bampi C, Jacquenet S, Lener D, Decimo D, Darlix JL 2004. The chaperoning and assistance roles of the HIV-1 nucleocapsid protein in proviral DNA synthesis and maintenance. Curr HIV Res 2: 79–92 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
