The identification of a new Giardia duodenalis assemblage in marine vertebrates and a preliminary analysis of G. duodenalis population biology in marine systems
- PMID: 20361967
- PMCID: PMC2900473
- DOI: 10.1016/j.ijpara.2010.02.015
The identification of a new Giardia duodenalis assemblage in marine vertebrates and a preliminary analysis of G. duodenalis population biology in marine systems
Abstract
Giardia duodenalis is an intestinal parasite of many vertebrates. The presence of G. duodenalis in the marine environment due to anthropogenic and wildlife activity is well documented, including the contributions from untreated sewage and storm water, agricultural run-off and droppings from terrestrial animals. Recently, studies have detected this protistan parasite in the faeces of marine vertebrates such as whales, dolphins, seals and shore birds. To explore the population biology of G. duodenalis in marine life, we determined the prevalence of G. duodenalis in two species of seal (Halichoerus grypus, Phoca vitulina vitulina and Phoca vitulina richardsi) from the east and west coasts of the USA, sequenced two loci from G. duodenalis-positive samples to assess molecular diversity and examined G. duodenalis distribution amongst these seals and other marine vertebrates along the east coast. We found a significant difference in the presence of G. duodenalis between east and west coast seal species. Only the zoonotic lineages of G. duodenalis, Assemblages A and B and a novel lineage, which we designated as Assemblage H, were identified in marine vertebrates. Assemblages A and B are broadly distributed geographically and show a lack of host specificity. Only grey seal (Halichoerus grypus) samples and one gull sample (Larus argentatus) from a northern location of Cape Cod, Massachusetts, USA, showed the presence of Assemblage H haplotypes; only one other study of harbour seals from the Puget Sound region of Washington, USA previously recorded the presence of an Assemblage H haplotype. Assemblage H sequences form a monophyletic clade that appears as divergent from the other seven Assemblages of G. duodenalis as these assemblages are from each other. The discovery of a previously uncharacterised lineage of G. duodenalis suggests that this parasite has more genetic diversity and perhaps a larger host range than previously believed.
Copyright 2010 Australian Society for Parasitology Inc. Published by Elsevier Ltd. All rights reserved.
Figures
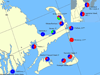


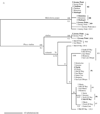


References
-
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J. Mol. Biol. 1990;215:403–410. - PubMed
-
- Cacciò SM, Ryan U. Molecular epidemiology of giardiasis. Mol. Biochem. Parasitol. 2008;160:75–80. - PubMed
-
- Cacciò SM, Sprong H. Giardia duodenalis: genetic recombination and its implications for taxonomy and molecular epidemiology. Exp. Parasitol. 2010;124:107–112. - PubMed
-
- Colegrove K, Greig DJ, Gulland FMD. Causes of live stranding of Northern Elephant Seals (Mirounga angustirostris) and Pacific Harbor Seals (Phoca vitulina) along the Central California Coast, 1992–2001. Aquatic mammals. 2005;31:1–10.
-
- Cooper M, Adam RD, Worobey M, Sterling CR. Population genetics provides evidence of recombination in Giardia. Curr. Biol. 2007;17:1–5. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

